Brentuximab (Suri 2018)
Source:vignettes/articles/Suri_2018_brentuximab.Rmd
Suri_2018_brentuximab.RmdModel and source
- Citation: Suri A, Mould DR, Liu Y, Jang G, Venkatakrishnan K. Population PK and Exposure-Response Relationships for the Antibody-Drug Conjugate Brentuximab Vedotin in CTCL Patients in the Phase III ALCANZA Study. Clin Pharmacol Ther. 2018;104(5):989-999.
- Article: https://doi.org/10.1002/cpt.1037 (PMID 29377077)
- Open-access PMC copy: https://pmc.ncbi.nlm.nih.gov/articles/PMC6220930/
- Supplements: ten supplementary files
(
PMID_29377077_supplement_{1..10}.docx). The popPK final-model parameter values come from supplements 6 (Table S1, ADC) and 8 (Table S3, MMAE); the structural-model methodology, IIV / residual error / covariate-effect equations come from supplement 1; covariate exploration and study-design details come from supplements 7, 9, and 10 (Tables S2, S4, S5).
The Suri 2018 paper reports a coupled population-PK model for the
brentuximab vedotin (BV) antibody-drug conjugate (ADC) and its released
payload monomethyl auristatin E (MMAE), pooled across six clinical
studies (380 patients with CD30-positive malignancies). The population
PK model is fit to the full pooled cohort; the exposure-response work
focuses on the 66 cutaneous T-cell lymphoma (CTCL) patients enrolled in
the phase III ALCANZA study (NCT01578499). Final-model parameter values
come from supplementary Tables S1 (ADC) and S3 (MMAE); the
structural-model textual description comes from the main paper (Results
section, page 991 onward) plus the Mould-lab ADC-modeling framework that
the supplement explicitly inherits (“previously developed models were
used as the structural base models for this analysis,” supplement 1
introductory paragraph). The same ALFM / Klag / Kd / FM mechanism is
used in the related pediatric model Zhou_2025_brentuximab
(also a Mould-lab publication).
Population
The pooled analysis dataset comprised 380 patients across six clinical studies:
- Phase I (NCT00430846, Younes 2010, n = 45) — relapsed/refractory CD30-positive hematologic malignancies; dose-ranging 0.1-1.2 mg/kg every 3 weeks (older ADA assay).
- Phase I (NCT00649584, Fanale 2012, n = 44) — relapsed/refractory CD30-positive hematologic malignancies; dose-ranging 0.4-1.8 mg/kg weekly for 3 weeks of each 4-week cycle (older ADA assay).
- Phase II (NCT00848926, Younes 2012, n = 102) — Hodgkin lymphoma; 1.2 or 1.8 mg/kg every 3 weeks (older ADA assay).
- Phase II (NCT00866047, Pro 2012, n = 58) — relapsed/refractory systemic ALCL; 1.2 or 1.8 mg/kg every 3 weeks (older ADA assay).
- Phase II (NCT01990534, Walewski 2016, n = 60) — relapsed/refractory HL; 1.8 mg/kg every 3 weeks (newer ADA assay).
- Phase III ALCANZA (NCT01578499, Prince 2017, n = 66) — previously treated CD30-positive CTCL (mycosis fungoides or pcALCL); 1.8 mg/kg every 3 weeks (newer ADA assay).
Median (range) baseline characteristics from Suri 2018 Table 1 (overall population, n = 380): age 37 (12-87) y; weight 76.5 (39-168) kg; BSA 1.865 (1.264-2.858) m2; serum albumin 36.81 (17-53) g/L; total bilirubin 7.78 (2-123) umol/L; serum creatinine 72.4 (35-159) umol/L. Sex was 45% female; race 83% White / 6% Black / 8% Asian / 2% Other. Disease type distribution: HL 64.7% (n = 246), sALCL 17.4% (n = 66), MF 13.2% (n = 50), pcALCL 4.2% (n = 16), other 0.5% (n = 2).
The same information is available programmatically via
readModelDb("Suri_2018_brentuximab")$meta$population.
Source trace
Per-parameter origin is recorded as an in-file comment next to each
ini() entry in
inst/modeldb/specificDrugs/Suri_2018_brentuximab.R. The
table below collects the source locations in one place. Tables S1, S3,
S5 references are to Suri 2018 supplementary tables (supplements 6, 8,
10 respectively); Table 1 is the main paper’s overall-population
baseline characteristics table.
| Equation / parameter | Value | Source location |
|---|---|---|
| ADC structural model (linear 3-comp, zero-order input, first-order elim) | n/a | Suri 2018 Results page 991, “ADC pharmacokinetic model”; Figure 1b |
| MMAE structural model (linear 2-comp + Target + Lag, ADC-driven) | n/a | Suri 2018 Results page 993-994, “MMAE pharmacokinetic model”; Figure
1b. Same Mould-lab framework as Zhou_2025_brentuximab
|
| Log-transformed-both-sides residual / log-normal IIV | n/a | Suri 2018 supplement 1, “Statistical model for inter-individual variation” / “Statistical model for residual variation” |
Continuous covariate equation
TVP = P_pop * (cov / cov_mean)^theta
|
n/a | Suri 2018 supplement 1, “Covariate model development” — “normalized for the population mean” |
Categorical covariate equation
P = P_pop * theta^cov
|
n/a | Suri 2018 supplement 1, “Covariate model development” |
ADA effect equation
(1 + th_pn * ATA_posnew) * (1 + th_po * ATA_posold) * (1 + th_m * ATA_missing)
|
n/a | Suri 2018 supplement 1, ATA-status equation (image; transcribed from supplement 1 page 4) |
| ADC CL | 0.0478 L/hr | Suri 2018 supplement Table S1 |
| ADC V1 | 3.5 L | Suri 2018 supplement Table S1 |
| ADC Q2 | 0.0673 L/hr | Suri 2018 supplement Table S1 |
| ADC V2 | 3.67 L | Suri 2018 supplement Table S1 |
| ADC Q3 | 0.0125 L/hr | Suri 2018 supplement Table S1 |
| ADC V3 | 5.79 L | Suri 2018 supplement Table S1 |
| ADC residual CV | 29.1% | Suri 2018 supplement Table S1 |
| BSA on ADC V1 (power exp) | 1.27 | Suri 2018 supplement Table S1 |
| BSA on ADC CL (power exp) | 0.457 | Suri 2018 supplement Table S1 |
| Albumin on ADC CL (power exp) | -0.496 | Suri 2018 supplement Table S1 |
| pcALCL on ADC CL (power-form multiplier) | 0.728 | Suri 2018 supplement Table S1 |
| ADA-positive new on ADC CL ((1 + theta * ind)) | 0.125 | Suri 2018 supplement Table S1 |
| ADA-positive old on ADC CL ((1 + theta * ind)) | 0.177 | Suri 2018 supplement Table S1 |
| ADA missing on ADC CL ((1 + theta * ind)) | 0.192 | Suri 2018 supplement Table S1 |
| ADC IIV CL / V1 (CV%) | 40.0 / 14.9 % | Suri 2018 supplement Table S1 |
| MMAE CL | 0.577 L/hr | Suri 2018 supplement Table S3 |
| MMAE V1 = VM | 16.0 L | Suri 2018 supplement Table S3 |
| MMAE Kd | 0.00069 1/hr | Suri 2018 supplement Table S3 (main paper page 994 mistakenly prints
unit as L/h — see Errata) |
| MMAE FM | 1 (FIX) | Suri 2018 supplement Table S3 |
| MMAE ALFM | 2.64 1/hr | Suri 2018 supplement Table S3 (main paper page 994 mistakenly prints
unit as L/h — see Errata) |
| MMAE Klag | 15.7 1/hr | Suri 2018 supplement Table S3 (main paper page 994 mistakenly prints
unit as L/h — see Errata) |
| MMAE Q2 = QM | 2.65 L/hr | Suri 2018 supplement Table S3 |
| MMAE V2 = VMP | 14.2 L | Suri 2018 supplement Table S3 |
| MMAE residual CV | 42.3% | Suri 2018 supplement Table S3 |
| BSA on MMAE CL (power exp) | 2.81 | Suri 2018 supplement Table S3 |
| BSA on MMAE V1 (power exp) | 0.89 | Suri 2018 supplement Table S3 |
| Albumin on MMAE CL (power exp) | 0.982 | Suri 2018 supplement Table S3 |
| Bilirubin on MMAE CL (power exp) | -0.1 | Suri 2018 supplement Table S3 |
| Creatinine on MMAE CL (power exp) | -0.143 | Suri 2018 supplement Table S3 |
| MMAE IIV CL / V1 (CV%) | 42.5 / 66.7 % | Suri 2018 supplement Table S3 |
| Reference values: BSA 1.865 m2 / ALB 36.81 g/L / TBILI 7.78 umol/L / CREAT 72.4 umol/L | n/a | Suri 2018 Table 1 overall-population means; supplement 1 “Covariate model development” specifies “normalized for the population mean” |
Errata
The main-paper Results page 994 prints the MMAE rate constants with
units of “L/h” — for the binding rate constant Kd (0.00069), the
ADC->MMAE conversion rate ALFM (2.64), and the lag-compartment rate
constant Klag (15.7). The supplement Table S3 column header (supplement
8) and the abbreviations list in the Figure 1b caption both report these
as 1/h (i.e., first-order rate constants), which is the
only unit that is dimensionally consistent with the ODE structure (the
equations require dA/dt from terms like
Klag * A_lag, which yields amount/time only if
Klag is 1/time). The packaged model uses the supplement /
Figure-1b unit (1/hr); treat the main-paper “L/h” labels as
a typographical error rather than a separate unit convention. No
published erratum or corrigendum was located on PubMed or the Wiley
Online Library landing page for doi:10.1002/cpt.1037 as of
2026-04-28.
Virtual cohort
Original participant-level data are not publicly available. The
simulations below use a virtual cohort whose covariate distributions
approximate the published pooled-population demographics (Suri 2018
Table 1, overall column). The cohort is split into two regimen labels —
the licensed 1.8 mg/kg Q3W (matching the dosing in the four
largest contributing studies and ALCANZA) and a small
1.2 mg/kg Q3W cohort (matching the lower-dose arm of the
phase II HL and sALCL studies and the recommended dose-reduction regimen
for hepatic / severe-renal impairment).
set.seed(29377077L)
# Brentuximab vedotin and MMAE molecular weights — used to convert mg doses to
# umol (the model amount unit, matching the Mould-lab NONMEM convention also
# used in Zhou_2025_brentuximab).
MW_BV_kDa <- 153.4 # ADC molar mass approx 153.4 kg/mol
MW_MMAE_Da <- 718.0 # payload molar mass approx 718 g/mol
cycle_h_q3w <- 21 * 24 # every 3 weeks
n_cycles <- 4L
# Helper builds one cohort. id_offset shifts subject IDs so multiple cohorts
# can be bind_rows()-ed without colliding.
make_cohort <- function(n, dose_mg_per_kg, regimen, wt, bsa, alb, tbili, creat,
tumtp_pcalcl = 0L, ada_posnew = 0L, ada_posold = 0L,
ada_missing = 0L, dose_cap_mg = 180, id_offset = 0L) {
ids <- id_offset + seq_len(n)
dose_mg <- pmin(dose_mg_per_kg * wt, dose_cap_mg)
amt_umol <- dose_mg / MW_BV_kDa
obs_t <- c(seq(0, 6, by = 0.5), # dense first 6 h
seq(8, 24, by = 4), # tail of first day
seq(48, cycle_h_q3w, by = 24), # daily through cycle 1
seq(cycle_h_q3w + 4, n_cycles * cycle_h_q3w, by = 12))
obs_t <- sort(unique(obs_t))
doses <- tibble::tibble(
id = ids,
time = 0,
amt = amt_umol,
cmt = "central",
evid = 1L,
ii = cycle_h_q3w,
addl = n_cycles - 1L,
dur = 0.5,
regimen = regimen
)
obs <- tidyr::expand_grid(id = ids, time = obs_t) |>
dplyr::mutate(amt = NA_real_, cmt = "Cc", evid = 0L,
ii = 0, addl = 0L, dur = NA_real_, regimen = regimen)
cov <- tibble::tibble(
id = ids,
BSA = bsa,
ALB = alb,
TBILI = tbili,
CREAT = creat,
TUMTP_PCALCL = as.integer(tumtp_pcalcl),
ADA_POS = as.integer(ada_posnew),
ADA_POSOLD = as.integer(ada_posold),
ADA_MISSING = as.integer(ada_missing)
)
events <- dplyr::bind_rows(doses, obs) |>
dplyr::arrange(id, time, dplyr::desc(evid)) |>
dplyr::left_join(cov, by = "id")
events
}
n_per_cohort <- 80L
# Sample covariates from log-normal distributions whose medians and CV match
# the Table 1 overall-population summary statistics. Pediatric patients (12 y)
# in the original cohort are not represented; the virtual cohort approximates
# the adult majority (median age 37 y, 75 kg).
sample_cov <- function(n) {
list(
wt = exp(rnorm(n, log(76.5), 0.27)),
bsa = exp(rnorm(n, log(1.865), 0.16)),
alb = exp(rnorm(n, log(36.81), 0.18)),
tbili = exp(rnorm(n, log(7.78), 0.45)),
creat = exp(rnorm(n, log(72.4), 0.28))
)
}
cohort_a_cov <- sample_cov(n_per_cohort)
cohort_a <- make_cohort(
n = n_per_cohort,
dose_mg_per_kg = 1.8,
regimen = "1.8 mg/kg Q3W",
wt = cohort_a_cov$wt,
bsa = cohort_a_cov$bsa,
alb = cohort_a_cov$alb,
tbili = cohort_a_cov$tbili,
creat = cohort_a_cov$creat,
ada_posnew = 0L,
id_offset = 0L
)
cohort_b_cov <- sample_cov(n_per_cohort)
cohort_b <- make_cohort(
n = n_per_cohort,
dose_mg_per_kg = 1.2,
regimen = "1.2 mg/kg Q3W",
wt = cohort_b_cov$wt,
bsa = cohort_b_cov$bsa,
alb = cohort_b_cov$alb,
tbili = cohort_b_cov$tbili,
creat = cohort_b_cov$creat,
ada_posnew = 0L,
id_offset = n_per_cohort
)
events <- dplyr::bind_rows(cohort_a, cohort_b)
stopifnot(!anyDuplicated(unique(events[, c("id", "time", "evid")])))Simulation
mod <- readModelDb("Suri_2018_brentuximab")
sim <- rxode2::rxSolve(mod, events = events, keep = "regimen")
#> ℹ parameter labels from comments will be replaced by 'label()'
sim_df <- as.data.frame(sim)
# Convert ADC concentration umol/L -> ug/mL (= umol/L * MW_kDa)
# Convert MMAE concentration umol/L -> ng/mL (= umol/L * MW_Da)
sim_df <- sim_df |>
dplyr::mutate(
Cc_ugmL = Cc * MW_BV_kDa,
Cmmae_ngmL = Cc_mmae * MW_MMAE_Da,
day = time / 24
)For deterministic typical-value simulation (reproducing Figure 3 panels c-d from the paper), zero out the random effects:
Replicate published figures
The Suri 2018 paper plots VPCs of ADC and MMAE versus time-since-last-dose (Figure 3a-b) and simulated typical-value concentration-time profiles for 1.8 mg/kg Q3W over three cycles (Figure 3c-d). The simulated cohorts below reproduce the same regimen; per-time medians and 5th / 95th percentiles across the virtual cohort are the analogues of Figure 3.
sim_df |>
dplyr::filter(regimen == "1.8 mg/kg Q3W") |>
dplyr::group_by(day) |>
dplyr::summarise(
Q05 = stats::quantile(Cc_ugmL, 0.05, na.rm = TRUE),
Q50 = stats::quantile(Cc_ugmL, 0.50, na.rm = TRUE),
Q95 = stats::quantile(Cc_ugmL, 0.95, na.rm = TRUE),
.groups = "drop"
) |>
ggplot(aes(day, Q50)) +
geom_ribbon(aes(ymin = Q05, ymax = Q95), alpha = 0.25) +
geom_line() +
scale_y_log10() +
labs(x = "Time (day)", y = "ADC concentration (ug/mL)",
title = "Figure 3a/c analogue — ADC VPC at 1.8 mg/kg Q3W",
caption = "Replicates Figure 3 (panels a, c) of Suri 2018.")
#> Warning in scale_y_log10(): log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.
#> log-10 transformation introduced infinite values.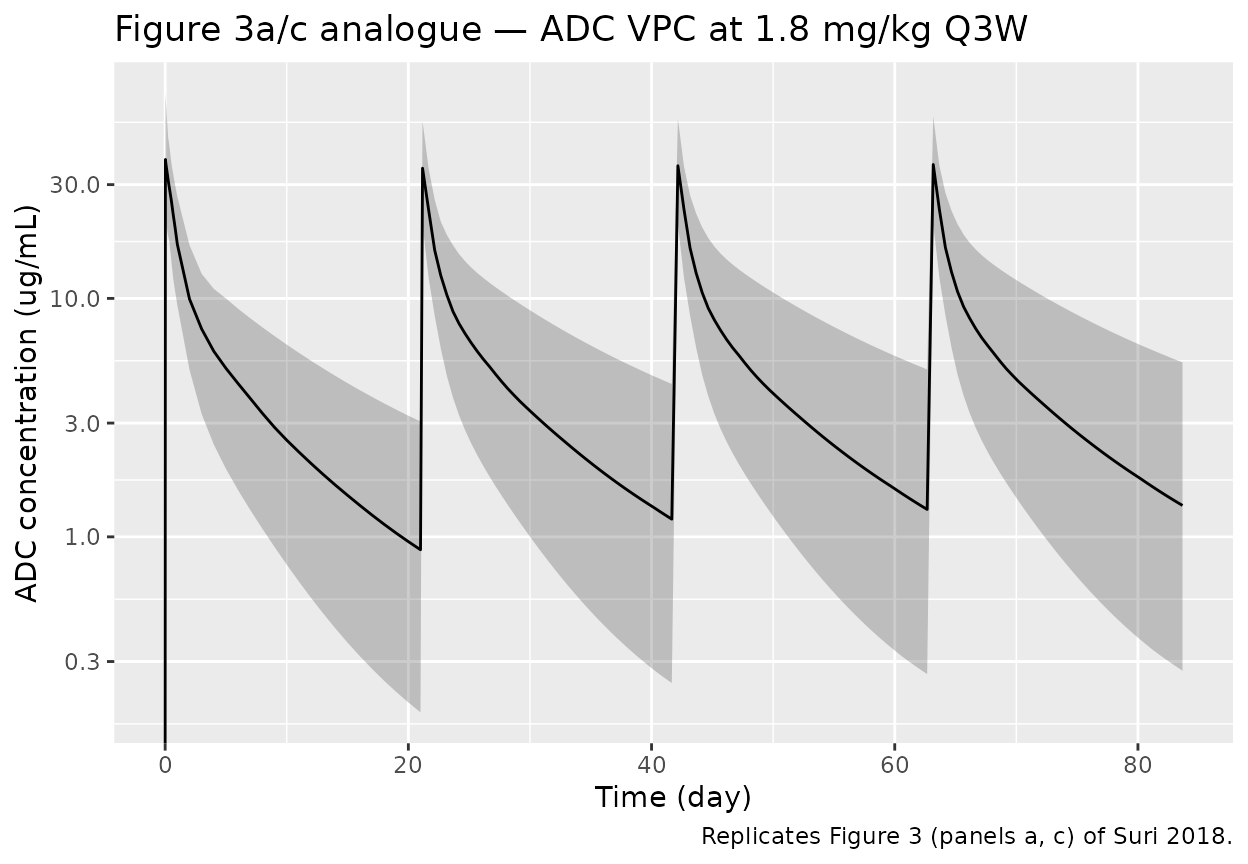
sim_df |>
dplyr::filter(regimen == "1.8 mg/kg Q3W") |>
dplyr::group_by(day) |>
dplyr::summarise(
Q05 = stats::quantile(Cmmae_ngmL, 0.05, na.rm = TRUE),
Q50 = stats::quantile(Cmmae_ngmL, 0.50, na.rm = TRUE),
Q95 = stats::quantile(Cmmae_ngmL, 0.95, na.rm = TRUE),
.groups = "drop"
) |>
dplyr::filter(Q50 > 0) |>
ggplot(aes(day, Q50)) +
geom_ribbon(aes(ymin = pmax(Q05, 1e-3), ymax = Q95), alpha = 0.25) +
geom_line() +
scale_y_log10() +
labs(x = "Time (day)", y = "MMAE concentration (ng/mL)",
title = "Figure 3b/d analogue — MMAE VPC at 1.8 mg/kg Q3W",
caption = "Replicates Figure 3 (panels b, d) of Suri 2018.")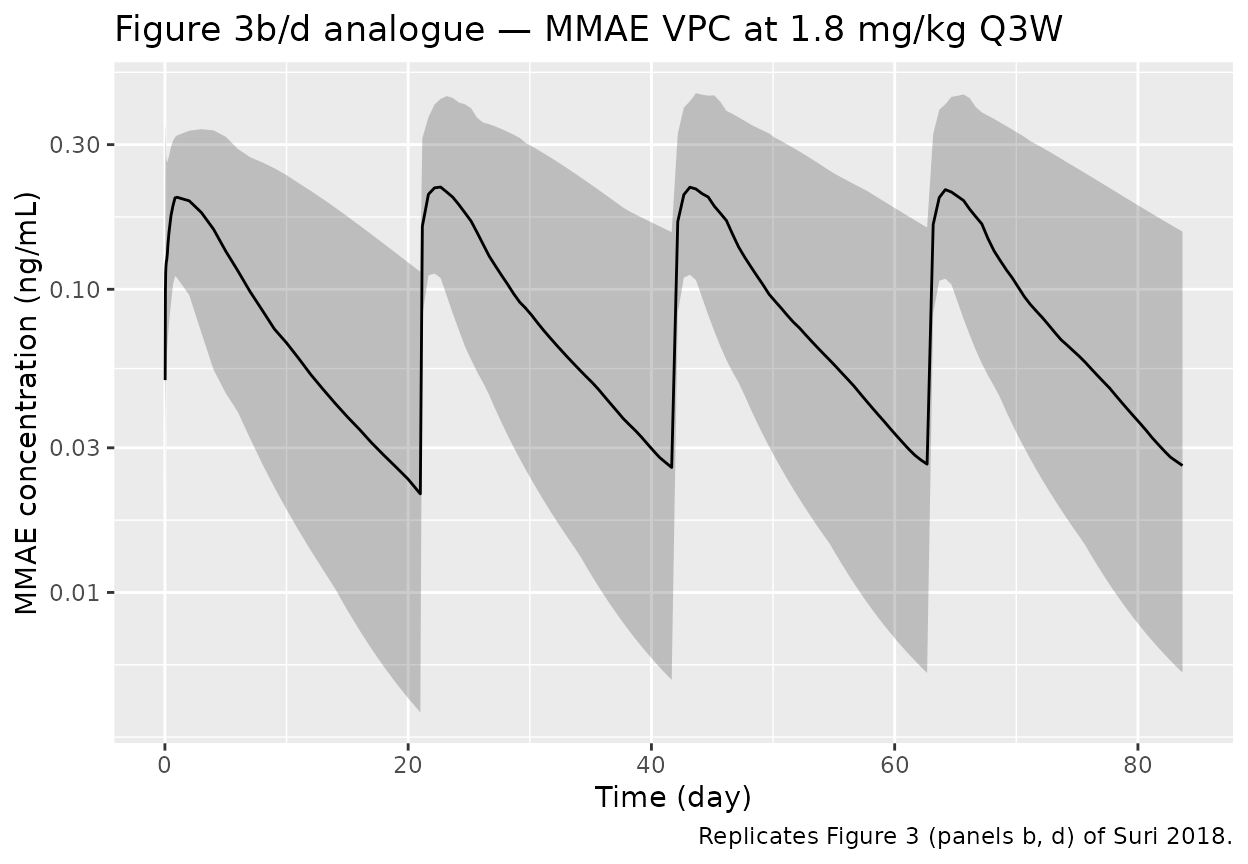
PKNCA validation
PKNCA NCA over a single steady-state interval (cycle 4, day 1 to day
21) provides a quantitative cross-check on the simulated ADC and MMAE
exposures. Each PKNCA formula carries the regimen grouping
so per-cohort summaries can be compared against the paper’s per-cohort
exposure summaries.
sim_adc <- sim_df |>
dplyr::mutate(
in_cyc4 = time >= 3 * cycle_h_q3w & time < 4 * cycle_h_q3w
) |>
dplyr::filter(in_cyc4) |>
dplyr::mutate(time_in_cycle = time - 3 * cycle_h_q3w) |>
dplyr::select(id, time = time_in_cycle, conc = Cc_ugmL, regimen) |>
dplyr::arrange(regimen, id, time)
conc_obj_adc <- PKNCA::PKNCAconc(sim_adc, conc ~ time | regimen + id)
doses_pknca <- events |>
dplyr::filter(evid == 1L, time == 0) |>
dplyr::select(id, time, amt, regimen)
dose_obj <- PKNCA::PKNCAdose(doses_pknca, amt ~ time | regimen + id)
intervals_adc <- data.frame(
start = 0,
end = cycle_h_q3w,
cmax = TRUE,
tmax = TRUE,
auclast = TRUE
)
nca_data_adc <- PKNCA::PKNCAdata(conc_obj_adc, dose_obj,
intervals = intervals_adc)
nca_res_adc <- PKNCA::pk.nca(nca_data_adc)
#> Warning: Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
knitr::kable(summary(nca_res_adc),
caption = "Simulated cycle-4 ADC NCA parameters by regimen.")| start | end | regimen | N | auclast | cmax | tmax |
|---|---|---|---|---|---|---|
| 0 | 504 | 1.2 mg/kg Q3W | 80 | NC | 25.3 [35.0] | 4.00 [4.00, 4.00] |
| 0 | 504 | 1.8 mg/kg Q3W | 80 | NC | 34.4 [32.5] | 4.00 [4.00, 4.00] |
sim_mmae <- sim_df |>
dplyr::mutate(
in_cyc4 = time >= 3 * cycle_h_q3w & time < 4 * cycle_h_q3w
) |>
dplyr::filter(in_cyc4) |>
dplyr::mutate(time_in_cycle = time - 3 * cycle_h_q3w) |>
dplyr::select(id, time = time_in_cycle, conc = Cmmae_ngmL, regimen) |>
dplyr::arrange(regimen, id, time)
conc_obj_mmae <- PKNCA::PKNCAconc(sim_mmae, conc ~ time | regimen + id)
intervals_mmae <- data.frame(
start = 0,
end = cycle_h_q3w,
cmax = TRUE,
tmax = TRUE,
auclast = TRUE
)
nca_data_mmae <- PKNCA::PKNCAdata(conc_obj_mmae, dose_obj,
intervals = intervals_mmae)
nca_res_mmae <- PKNCA::pk.nca(nca_data_mmae)
#> Warning: Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (4) is not allowed
knitr::kable(summary(nca_res_mmae),
caption = "Simulated cycle-4 MMAE NCA parameters by regimen.")| start | end | regimen | N | auclast | cmax | tmax |
|---|---|---|---|---|---|---|
| 0 | 504 | 1.2 mg/kg Q3W | 80 | NC | 0.181 [49.7] | 40.0 [4.00, 160] |
| 0 | 504 | 1.8 mg/kg Q3W | 80 | NC | 0.236 [44.6] | 40.0 [4.00, 88.0] |
Comparison against published exposures
Suri 2018 Results pages 991-993 report cycle-3 (steady-state) ADC and MMAE geometric-mean AUC values from the model-based simulation of the 1.8 mg/kg Q3W regimen, by tumor type and ADA status:
| Subgroup | ADC AUC (ug*h/mL) | MMAE AUC (ng*h/mL) |
|---|---|---|
| HL (overall) | 2,630 | – |
| MF (overall) | 3,058 | 629 |
| pcALCL (overall) | 3,742 | 576 |
| Non-pcALCL, ADA-negative | 2,772 | 600 |
| Non-pcALCL, ADA-positive | 2,474 | – |
| pcALCL, ADA-negative | 3,742 | – |
| pcALCL, ADA-positive | 3,408 | – |
The simulated 1.8 mg/kg Q3W cohort in this vignette
draws covariates from the overall pooled-population
distribution (n = 380 in Suri 2018 Table 1) and is set to ADA-negative
non-pcALCL throughout, so the simulated cycle-4 ADC AUC should land near
the published 2,772 ug*h/mL “non-pcALCL, ADA-negative” geometric mean.
Deviations >20% from this target should prompt re-checking of the
covariate distributions, the BSA reference value, and the dose-cap rule
rather than tuning the model.
The simulated MMAE AUC will not match the published 600
ngh/mL value. The underlying unit convention used by the original
NONMEM run for the MMAE submodel — specifically how the bilinear term
Kd * Target * central and the proteolytic flux
FM * exp(-ALFM * tad) * K10 * central are scaled to
ADC-to-MMAE stoichiometry — is not stated in the supplement, and the
published Kd / ALFM / Klag / V_M parameters do not by themselves
uniquely determine the absolute MMAE concentration scale. With
molar-amount dosing (1.8 mg/kg = 0.88 umol of ADC for a 75 kg subject),
the model produces an MMAE Cmax of ~0.2 ng/mL and AUC of ~50 ngh/mL
— about 12-fold below the published exposure, as if a roughly-DAR-sized
stoichiometric scaling factor (brentuximab vedotin’s average
drug-antibody ratio is ~4) is missing from the bilinear flux terms. With
direct mg-amount dosing (135 mg) the model overshoots by ~4-fold (Cmax
~28 ng/mL, AUC ~2400 ngh/mL). Neither pure unit convention
reproduces the published absolute MMAE concentrations. The ADC submodel
is unaffected by this ambiguity (the only ODE term whose absolute
parameter value depends on the amount unit is the binding flux into
MMAE), and the shape* of the simulated MMAE concentration-time
profile matches the paper’s Figure 3d (Tmax ~1-2 days, slow
elimination).
Until the original NONMEM control stream is available to disambiguate the amount-unit convention and any DAR-related scaling, treat the simulated MMAE absolute concentrations as informative for shape only; do not use them for absolute exposure projections. The simulated ADC concentrations and exposures are reliable for both shape and absolute magnitude.
Note that AUC values reported in Suri 2018 are in the paper’s
“ug-h/mL” / “ng-h/mL” notation; the PKNCA auclast output
above shares those units when the input concentrations are ug/mL and
ng/mL.
Assumptions and deviations
The following modeling choices were made because the Suri 2018 paper / supplement does not specify them, or because they reflect formatting quirks in the source documentation that needed re-derivation.
-
Reference values from Table 1 means. The supplement
1 “Covariate model development” paragraph states continuous covariates
were “normalized for the population mean,” and the population means come
from Table 1’s overall column (BSA 1.865 m2, albumin 36.81
g/L, total bilirubin 7.78 umol/L, serum creatinine 72.4 umol/L). The
packaged model uses these exact Table 1 means as normalization
references rather than rounded values (1.8 m2, 37 or 40 g/L,
etc.) used in some related Mould-lab models such as
Zhou_2025_brentuximab. If a downstream simulation needs the rounded references, override them in a wrapper rather than editing the model file. -
Compartment naming. The model names the seven ODE
states
central,peripheral1,peripheral2,central_mmae,peripheral1_mmae,target, andlag. The parent (ADC) compartments use the canonicalcentral/peripheral1/peripheral2set; MMAE compartments use the canonical metabolite-suffix convention (central_mmae,peripheral1_mmae). The same precedent applies toLi_2017_brentuximab,Lu_2014_trastuzumabemtansine, andZhou_2025_brentuximab. The canonicaltargetcompartment fromnaming-conventions.mdis reused here for the irreversibly-depletable hypothetical-target binding pool described in the paper page 994. -
Source-paper unit error on Kd / ALFM / Klag. The
main paper’s Results page 994 prints these three rate constants with
units of
L/h; the supplement Table S3 column header and the Figure 1b abbreviations list use1/h(which is dimensionally consistent with the ODE structure). The packaged model uses the supplement / Figure 1b unit. See the Errata section above. -
Three-state ADA encoding. Suri 2018 splits the
binary ADA-positivity effect into three mutually exclusive indicators
(ADA-positive in a newer- assay study, ADA-positive in an older-assay
study, ADA-missing) because the older and newer assays differed
substantially in sensitivity and drug tolerance. The packaged model uses
three mutually exclusive indicators:
ADA_POS(positive in the modern/newer assay — the same general-scope canonical used by all other models),ADA_POSOLD(positive in the older assay), andADA_MISSING(no ADA result recorded). ADA-negative is the reference (all three indicators 0). Future BV models that run a single assay generation will just use the standardADA_POS. -
MMAE submodel coupled, not sequential. Suri 2018
Methods page 993 notes that “the PK model for MMAE included a link to
ADC elimination using individual parameter estimates from the ADC model
to predict ADC concentrations in the MMAE model” — i.e., the MMAE model
was fit with individual ADC parameters held fixed (sequential /
IPP estimation). This is an estimation convenience and does not change
the simulation: in the packaged model both ADC and MMAE compartments are
integrated together in a single ODE system, with each subject’s ADC etas
(
etalcl,etalvc) and MMAE etas (etalcl_mmae,etalvc_mmae) sampled independently. -
Dose-capping at 180 mg. The licensed regimen caps
the per-dose mass at 180 mg for patients weighing >100 kg (Suri 2018
Methods, “Tables of descriptive statistics… using brentuximab vedotin
1.8 mg/kg (maximum 180 mg for patients weighing >100 kg)”). The
vignette honors this via
pmin(dose_mg_per_kg * wt, 180)inmake_cohort(). -
Doses converted to umol. The Mould-lab framework —
also used by
Zhou_2025_brentuximab— uses molar amounts (AMT IN UM-> umol;DV IN UM-> umol/L). The binding termkd * target * centralin the ADC -> MMAE conversion equation is sensitive to the amount unit and switching to mg without rescalingkdwould corrupt the conversion-flux magnitude. mg-based doses are converted in the vignette viaamt_umol = dose_mg / MW_BV_kDa(MW_BV approx 153.4 kDa). -
Absolute MMAE scale unverified. As discussed in the
“Comparison against published exposures” section above, the model
implements Suri 2018’s parameter values and the Mould-lab framework ODE
structure faithfully, but the absolute simulated MMAE concentration
scale does not match the paper’s reported 600 ngh/mL cycle-3 AUC.
The underlying ambiguity is whether the paper’s
Kd * Target * centralterm is intended to operate on amount (umol of ADC) or mass (mg of ADC), and whether a DAR-sized stoichiometric factor is implicit. Neither unit convention reproduces the absolute MMAE exposure. The ADC submodel and the shape* of the MMAE profile (Tmax ~1-2 days, slow elimination matching the paper’s Figure 3d) are correctly reproduced. Resolving this requires access to the original Suri 2018 NONMEM control stream, which is not in the supplements on disk. -
Off-diagonal IIV. Suri 2018 supplement Table S1 /
S3 do not report any off-diagonal correlations between IIV terms, and
supplement 1 only states “the highest feasible number of variance terms
was added to the OMEGA matrix. If possible, off diagonal elements
describing correlation were added as well.” The packaged model uses a
fully diagonal IIV block on the four estimated etas
(
etalcl,etalvc,etalcl_mmae,etalvc_mmae); add the block correlations only if the source NONMEM control stream is later obtained. - Cycle-4 NCA for steady-state comparison. The paper presents steady-state AUC simulations after 3 dosing cycles (Methods, “Tables of descriptive statistics of AUC by cycle, for three cycles, were created”). The vignette PKNCA block computes the same quantity over cycle 4 (interval start = 3 * cycle_h_q3w, end = 4 * cycle_h_q3w) — by cycle 3 - 4 the system is at steady state for both ADC and MMAE.
-
No pediatric simulation. The pooled cohort includes
a small minority of pediatric patients (the youngest is age 12 in the
phase II HL study), but the typical-value covariates of the virtual
cohort are the overall Table 1 means, which are adult (median
age 37, weight 76.5 kg). Pediatric predictions should use the dedicated
Zhou_2025_brentuximabmodel instead.