Omalizumab (Hayashi 2007)
Source:vignettes/articles/Hayashi_2007_omalizumab.Rmd
Hayashi_2007_omalizumab.Rmd
library(nlmixr2lib)
library(rxode2)
#> rxode2 5.1.2 using 2 threads (see ?getRxThreads)
#> no cache: create with `rxCreateCache()`
library(dplyr)
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, union
library(tidyr)
library(ggplot2)
library(PKNCA)
#>
#> Attaching package: 'PKNCA'
#> The following object is masked from 'package:stats':
#>
#> filterOmalizumab–IgE binding population PK/PD model
Omalizumab is a humanized anti-IgE IgG1 monoclonal antibody used in moderate-to-severe atopic asthma. It interrupts the allergic cascade by binding free serum IgE, lowering its concentration, and downmodulating FcεRI on basophils and mast cells. Hayashi et al. (2007) developed a mechanism-based population PK/PD model in 202 Japanese subjects (single- and multiple-dose studies 1101 and 1305) that describes three serum entities — free omalizumab, free IgE, and the omalizumab–IgE complex — each with their own clearance and volume of distribution, coupled by an instantaneous-equilibrium binding step (law of mass action) with a concentration-dependent dissociation constant. Body weight modifies omalizumab CL and Vd; baseline IgE modifies IgE CL and IgE production rate. The model was externally validated against 531 White patients (studies 007/008/009).
- Citation: Hayashi N, Tsukamoto Y, Sallas WM, Lowe PJ. A mechanism-based binding model for the population pharmacokinetics and pharmacodynamics of omalizumab. Br J Clin Pharmacol. 2007;63(5):548-561. doi:10.1111/j.1365-2125.2006.02803.x (PMID 17096680).
- Article: https://doi.org/10.1111/j.1365-2125.2006.02803.x
- PMID: 17096680
Population
The model-building dataset comprises 202 Japanese subjects from two clinical studies (Hayashi 2007 Table 1):
| Study | Indication | n (used) | Design | Body weight (kg) | Baseline IgE (ng/mL) |
|---|---|---|---|---|---|
| 1101 | Healthy atopic volunteers | 48 | Single SC dose 75–375 mg | 62.5 ± 6.4 (51–79) | 811 ± 473 (204–2143) |
| 1305 | Seasonal allergic rhinitis (Japan) | 154 | Multiple SC dosing per the body-weight × IgE table (Table 2) | 60.5 ± 10.2 (42–101) | 373 ± 317 (53–1316) |
A total of 3192 observation records (1037 omalizumab, 1191 total IgE,
964 free IgE) were included; 275 free-IgE values above the 150 ng/mL
upper limit of quantification were excluded. The same metadata is
available programmatically through
readModelDb("Hayashi_2007_omalizumab")$population.
Source trace
Per-parameter origin is recorded as in-file comments next to each
ini() entry in
inst/modeldb/specificDrugs/Hayashi_2007_omalizumab.R. The
table collects them for review.
| Equation / parameter | Value (paper) | Value (file) | Source |
|---|---|---|---|
lka (SC absorption rate) |
0.0200 1/h | log(0.0200·24) 1/d | Table 3 |
lcl (apparent CL of free omalizumab at 61.1 kg) |
7.32 mL/h | log(0.00732·24) L/d | Table 3 |
ldcl_complex (apparent excess CL of complex) |
5.86 mL/h | log(0.00586·24) L/d | Table 3 |
lcl_ige (apparent CL of free IgE at 482.4 ng/mL) |
71.0 mL/h | log(0.071·24) L/d | Table 3 |
lvc (apparent V of omalizumab and IgE at 61.1 kg) |
5900 mL | log(5.9) L | Table 3 (V_E/f = V_X/f, footnote ‡) |
lvc_complex (apparent V of complex) |
3630 mL | log(3.63) L | Table 3 |
lp_ige (apparent IgE production rate at 482.4
ng/mL) |
30.3 µg/h (= 0.1595 nmol/h) | log(0.1595·24) nmol/d | Table 3, footnote † |
lkd0 (Kd at X_TX = X_TE) |
1.07 nM | log(1.07) nM | Table 3 |
e_wt_cl (BW exponent on CL_X) |
0.911 | 0.911 | Table 3 |
e_wt_vc (BW exponent on V_X) |
0.658 | 0.658 | Table 3 |
e_ige_cl_ige (IgE0 exponent on CL_E) |
−0.281 | −0.281 | Table 3 |
e_ige_p_ige (IgE0 exponent on P_E) |
0.657 | 0.657 | Table 3 |
alpha (concentration dependence on Kd) |
0.157 | 0.157 | Table 3 |
etalka |
CV 39.9% | ω² = 0.14773 | Table 3 |
etalcl |
CV 20.3% | ω² = 0.04042 | Table 3 |
etaldcl_complex |
CV 34.9% | ω² = 0.11488 | Table 3 |
etalvc (shared by V_X and V_E) |
CV 13.0% | ω² = 0.01679 | Table 3 |
etalvc_complex |
CV 25.0% | ω² = 0.06062 | Table 3 |
etalcl_ige + etalp_ige (block; r = 0.968) |
CV 25.3% / 23.1% | ω² = 0.06205, cov 0.05496, ω² = 0.05197 | Table 3 |
propSd / propSd_totalIgE /
propSd_freeIgE (Y = F·exp(ε)) |
σ ≈ 16.7% / 21.1% / 21.8% | 0.167 / 0.211 / 0.218 | Table 3 |
| Eq. dX_SC/dt = −ka·X_SC + f·D·δ(t−tD) | n/a | n/a | Methods, page 552 |
| Eq. dX_TX/dt = ka·X_SC − CL_X·C_fX − CL_C·C_C | n/a | n/a | Methods, page 552 |
| Eq. dX_TE/dt = P_E − CL_E·C_fE − CL_C·C_C | n/a | n/a | Methods, page 552 |
| Quadratic X_C = ½{S − √(S² − 4·X_TX·X_TE)}, S = X_TX + X_TE + Kd·V_X·V_E/V_C | n/a | n/a | Methods, page 552 |
| Concentration-dependent Kd = Kd0·(X_TX/X_TE)^α | n/a | n/a | “The dissociation constant”, page 552 |
| Power covariates: CL_X, V_X on (WT/61.1); CL_E, P_E on (IgE0/482.4) | n/a | n/a | Page 555 equations |
| MW(omalizumab) = 150 kDa; MW(IgE) = 190 kDa | 150 / 190 kDa | 150 / 190 kDa | Methods, page 552 |
| Subcutaneous bioavailability used to translate apparent values to absolute | f = 0.62 (ref [28]) | not encoded; CL/f and V/f in ini()
|
Discussion, page 559 |
Virtual cohort
Original observed data are not publicly available. The cohort below reproduces the four single-dose groups of study 1101 (75, 150, 300, and 375 mg SC) — the only design with a dense sampling schedule (Hayashi 2007 Methods, page 549). Body weight and baseline IgE are sampled from the study-1101 distribution (mean 62.5 kg ± 6.4 kg; mean 811 ng/mL ± 473 ng/mL, log-normal).
set.seed(20070501L) # paper publication month YYYYMM01
study_1101_doses <- c(75, 150, 300, 375)
n_per_dose <- 12L
make_cohort <- function(n, dose_mg, id_offset = 0L) {
data.frame(
id = id_offset + seq_len(n),
dose = dose_mg,
WT = pmax(45, pmin(80, rnorm(n, mean = 62.5, sd = 6.4))),
IGE = pmax(200, pmin(2200,
exp(rnorm(n, mean = log(811) - 0.5 * log(1 + (473 / 811)^2),
sd = sqrt(log(1 + (473 / 811)^2))))))
)
}
cohorts <- bind_rows(lapply(seq_along(study_1101_doses), function(i) {
make_cohort(n_per_dose, study_1101_doses[i],
id_offset = (i - 1L) * n_per_dose)
}))
obs_times <- c(0, 0.5, 1, 2, 4, 7, 10, 14, 28, 42, 56, 70, 84) # days, study 1101 schedule
events <- cohorts |>
rowwise() |>
do({
row <- .
et_obj <- rxode2::et(time = 0, amt = row$dose, cmt = "depot") |>
rxode2::et(obs_times, cmt = "Cc")
et_obj$id <- row$id
et_obj$WT <- row$WT
et_obj$IGE <- row$IGE
et_obj$dose <- row$dose
et_obj
}) |>
ungroup() |>
as.data.frame()
stopifnot(!anyDuplicated(unique(events[, c("id", "time", "evid")])))Simulation
mod <- readModelDb("Hayashi_2007_omalizumab")
sim <- rxode2::rxSolve(mod, events = events,
keep = c("dose", "WT", "IGE"),
returnType = "data.frame", addDosing = FALSE)
#> ℹ parameter labels from comments will be replaced by 'label()'
sim$dose_label <- factor(paste0(sim$dose, " mg"),
levels = paste0(study_1101_doses, " mg"))Replicate published Figure 3 — single-dose profiles by dose level
Hayashi 2007 Figure 3 shows individual observed serum concentrations of omalizumab, total IgE, and free IgE in study 1101 over 84 days, with 80% prediction intervals. We reproduce the cohort means with 80% prediction intervals from the simulated population.
sim |>
group_by(time, dose_label) |>
summarise(
Q10 = quantile(Cc, 0.10, na.rm = TRUE),
Q50 = quantile(Cc, 0.50, na.rm = TRUE),
Q90 = quantile(Cc, 0.90, na.rm = TRUE),
.groups = "drop"
) |>
ggplot(aes(time, Q50)) +
geom_ribbon(aes(ymin = Q10, ymax = Q90), alpha = 0.25) +
geom_line() +
facet_wrap(~ dose_label, scales = "free_y") +
labs(x = "Time (days)", y = "Total omalizumab (µg/mL)",
title = "Hayashi 2007 Figure 3, omalizumab panel",
caption = "Lines: median; ribbon: 10th–90th percentile from 12 simulated subjects per dose.")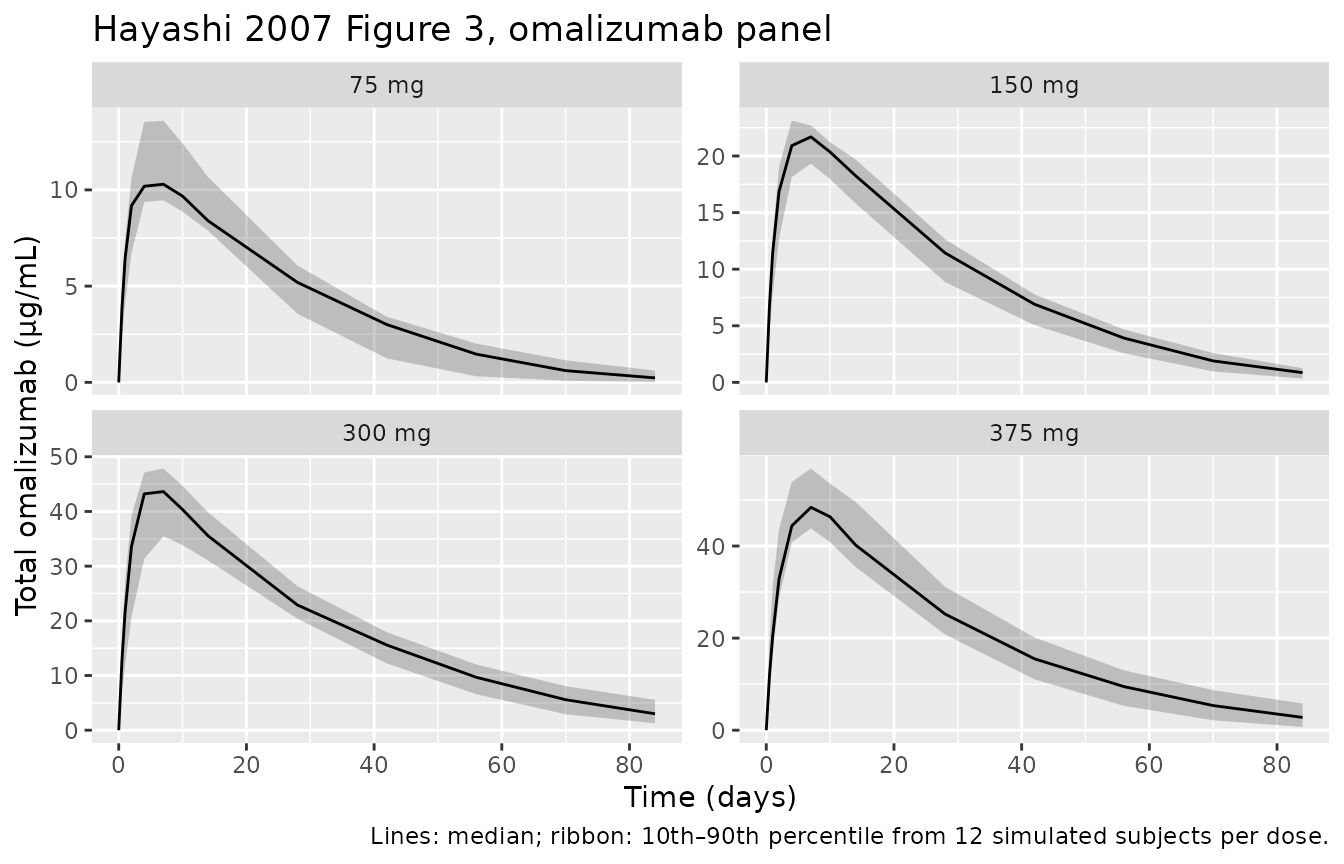
Replicates Figure 3 of Hayashi 2007 (top row, omalizumab vs time by single-dose group, study 1101).
sim |>
group_by(time, dose_label) |>
summarise(
Q10 = quantile(freeIgE, 0.10, na.rm = TRUE),
Q50 = quantile(freeIgE, 0.50, na.rm = TRUE),
Q90 = quantile(freeIgE, 0.90, na.rm = TRUE),
.groups = "drop"
) |>
ggplot(aes(time, Q50)) +
geom_ribbon(aes(ymin = Q10, ymax = Q90), alpha = 0.25) +
geom_line() +
facet_wrap(~ dose_label) +
scale_y_log10() +
geom_hline(yintercept = 12, lty = 2, colour = "grey40") +
geom_hline(yintercept = 21, lty = 2, colour = "grey40") +
labs(x = "Time (days)", y = "Free IgE (ng/mL, log scale)",
title = "Hayashi 2007 Figure 3, free-IgE panel",
caption = "Dashed grey lines indicate the 12–21 ng/mL clinical-efficacy band cited in Hayashi 2007.")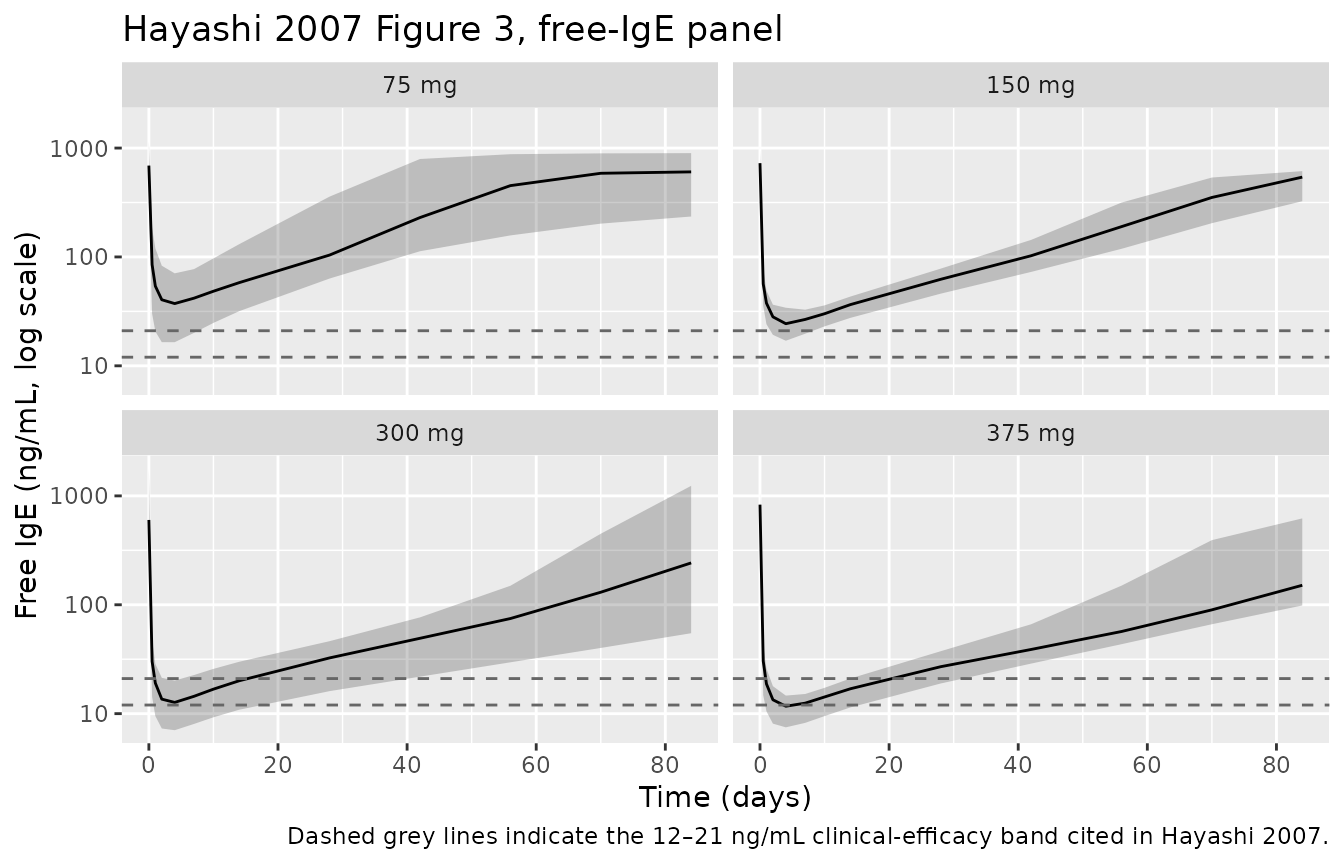
Replicates Figure 3 of Hayashi 2007 (free IgE vs time by single-dose group, study 1101).
sim |>
group_by(time, dose_label) |>
summarise(
Q10 = quantile(totalIgE, 0.10, na.rm = TRUE),
Q50 = quantile(totalIgE, 0.50, na.rm = TRUE),
Q90 = quantile(totalIgE, 0.90, na.rm = TRUE),
.groups = "drop"
) |>
ggplot(aes(time, Q50)) +
geom_ribbon(aes(ymin = Q10, ymax = Q90), alpha = 0.25) +
geom_line() +
facet_wrap(~ dose_label) +
labs(x = "Time (days)", y = "Total IgE (ng/mL)",
title = "Hayashi 2007 Figure 3, total-IgE panel")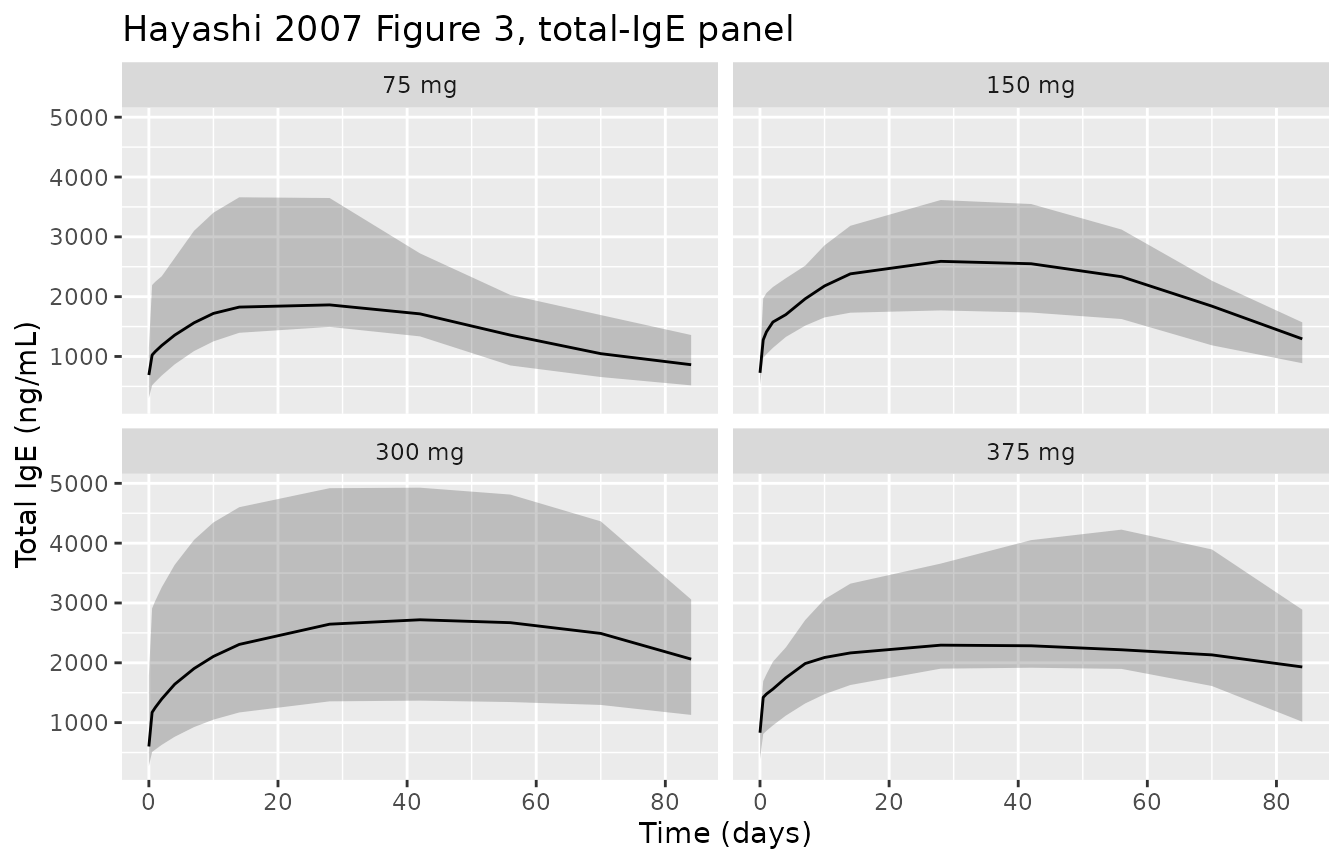
Replicates Figure 3 of Hayashi 2007 (total IgE vs time by single-dose group, study 1101).
PKNCA validation — omalizumab
PKNCA computes Cmax, Tmax, AUC(0,∞), and terminal half-life on the simulated total-omalizumab profile. Hayashi 2007 reports a noncompartmental terminal half-life of 18.2 days for total omalizumab in study 1101 (Discussion, page 558) and a model-derived half-life of 23 days for free omalizumab. The compartmental Cc output is total omalizumab (free + complex), so the NCA t½ is expected to land between those two values.
sim_nca <- sim |>
filter(!is.na(Cc)) |>
select(id, time, Cc, dose_label)
dose_df <- events |>
filter(evid == 1) |>
mutate(dose_label = factor(paste0(amt, " mg"),
levels = paste0(study_1101_doses, " mg"))) |>
select(id, time, amt, dose_label)
conc_obj <- PKNCA::PKNCAconc(sim_nca, Cc ~ time | dose_label + id)
dose_obj <- PKNCA::PKNCAdose(dose_df, amt ~ time | dose_label + id)
intervals <- data.frame(
start = 0,
end = Inf,
cmax = TRUE,
tmax = TRUE,
aucinf.obs = TRUE,
half.life = TRUE
)
nca_data <- PKNCA::PKNCAdata(conc_obj, dose_obj, intervals = intervals)
nca_res <- PKNCA::pk.nca(nca_data)
nca_summary <- summary(nca_res)
knitr::kable(nca_summary, caption = "Simulated NCA parameters by single-dose group (study 1101).")| start | end | dose_label | N | cmax | tmax | half.life | aucinf.obs |
|---|---|---|---|---|---|---|---|
| 0 | Inf | 75 mg | 12 | 10.7 [13.0] | 7.00 [4.00, 10.0] | 11.8 [5.12] | 332 [25.1] |
| 0 | Inf | 150 mg | 12 | 20.8 [16.1] | 7.00 [4.00, 10.0] | 14.8 [5.58] | 745 [18.4] |
| 0 | Inf | 300 mg | 12 | 44.6 [14.0] | 7.00 [4.00, 7.00] | 17.9 [4.42] | 1650 [13.8] |
| 0 | Inf | 375 mg | 12 | 49.9 [14.4] | 7.00 [4.00, 10.0] | 18.1 [4.87] | 1810 [22.4] |
Comparison against published values
Hayashi 2007 reports only a single global noncompartmental t½ for total omalizumab in study 1101 (18.2 days; Discussion, page 558) and the model-derived half-lives of free omalizumab (23 days) and IgE (2.4 days). The simulated terminal t½ for total omalizumab in the typical patient should sit between the two model-derived bounds (23 days for free omalizumab; faster for the complex due to ΔCL_C = 5.86 mL/h on top of CL_X = 7.32 mL/h). No per-dose NCA table is published, so a fully quantitative side-by-side comparison is not possible.
Mechanism check — typical-value pre-treatment steady state
Without drug, X_TE evolves under
dX_TE/dt = P_E − CL_E·C_fE. The model’s typical-value
pre-treatment steady-state free-IgE concentration is
(P_E / CL_E) · MW_IgE = 426.8 ng/mL. The covariate
reference value is 482.4 ng/mL — a small (~12%) intentional offset
because P_E and CL_E are independently estimated and not constrained to
reproduce IgE0 at steady state. With the model’s initial condition
X_TE(0) = (IGE / MW_IgE)·V_E, free IgE starts at the
observed baseline and drifts toward the model SS over the first few days
when there is no drug; once dosing begins the binding kinetics dominate
this small drift.
mod_typ <- rxode2::zeroRe(mod)
#> ℹ parameter labels from comments will be replaced by 'label()'
ev_nodose <- rxode2::et(seq(0, 30, by = 1), cmt = "Cc")
sim_nodose <- rxode2::rxSolve(mod_typ, events = ev_nodose,
params = c(WT = 61.1, IGE = 482.4),
returnType = "data.frame", addDosing = FALSE)
#> ℹ omega/sigma items treated as zero: 'etalka', 'etalcl', 'etaldcl_complex', 'etalvc', 'etalvc_complex', 'etalcl_ige', 'etalp_ige'
ggplot(sim_nodose, aes(time, freeIgE)) +
geom_line() +
geom_hline(yintercept = 482.4, lty = 2, colour = "grey40") +
geom_hline(yintercept = (0.1595 / 0.071) * 190, lty = 3, colour = "grey40") +
labs(x = "Time (days, no drug)", y = "Free IgE (ng/mL)",
title = "Pre-treatment steady-state drift toward the model SS",
caption = "Dashed: observed IgE0 = 482.4 ng/mL (initial). Dotted: model SS = P_E/CL_E·MW_IgE ≈ 426.8 ng/mL.")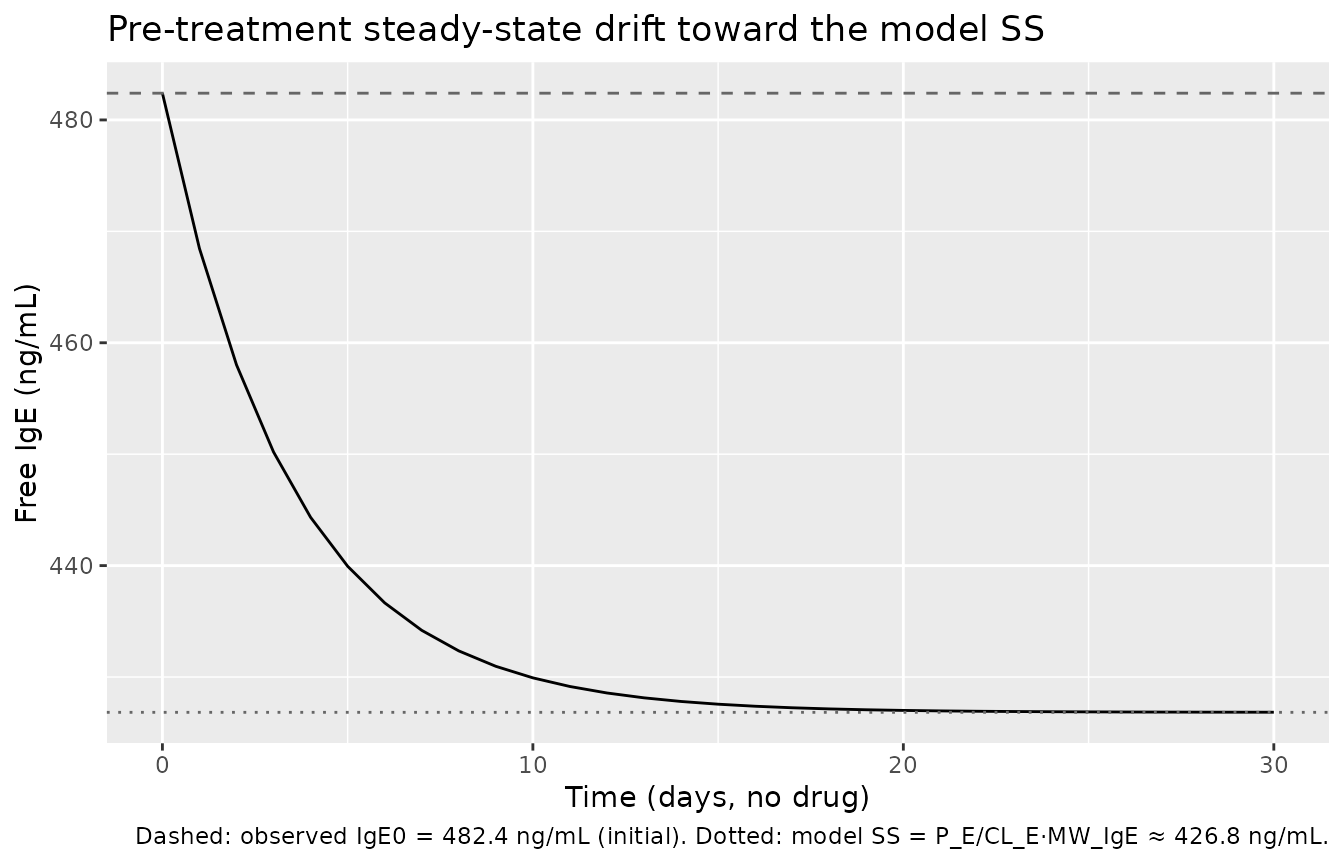
Assumptions and deviations
- Bioavailability not encoded explicitly. The paper reports apparent parameters (CL_X/f, V_X/f, etc.) and notes f = 0.62 from a separate source (Discussion, page 559). The model file uses the apparent values directly with no f scaling on the dose, which faithfully reproduces the predicted measured concentrations in the original NONMEM fit. Absolute clearances and volumes can be recovered by multiplying the apparent values by 0.62.
-
Time unit conversion h → d. The paper reports CL
and ka in 1/h, consistent with NONMEM convention. The model file
converts to 1/day by multiplying each rate by 24, with the conversion
shown in the
ini()source-trace comment for every rate parameter. Half-lives derived from the converted values match the paper (23 days for free omalizumab, 2.4 days for IgE). -
Residual error written as proportional rather than exact
log-normal. Hayashi 2007 uses Y = F · exp(ε); for σ in the
0.17–0.22 range encountered here, Y ≈ F·(1 + ε), so a proportional error
model with
propSd = σis faithful to within third-order ε terms. nlmixr2’sprop()keyword maps to this approximation; the alternativelnorm()parameterization could be used in future revisions. -
V_E/f shared with V_X/f. Footnote ‡ of Table 3
states V_E/f was fixed equal to V_X/f because separate estimation did
not converge. The model file enforces this by setting
v_ige <- vc(same individual parameter, including the sameetalvcIIV). -
Concentration-dependent Kd at t = 0. With
central(paper X_TX) = 0 at t = 0, Kd = Kd0·(0/total_target)^α = 0 for α > 0, S = total_target, and the negative root yields X_C = 0. This matches the physical expectation (no drug, no complex). After dosing beginscentralbecomes strictly positive and the expression evaluates without numerical issues. -
Compartment names. The paper’s X_TX (total
omalizumab amount in serum, free + complex) maps to the canonical
centralcompartment; X_TE (total IgE amount, free + complex) maps tototal_target. This follows the canonical TMDD compartment names registered inreferences/naming-conventions.mdfor QSS-style total-amount parameterizations (Gibiansky 2008). Thecomplexspecies is algebraically derived from the equilibrium relation rather than carried as a separate ODE state. - External validation cohort not modelled. The paper validates predictions against 531 White patients in studies 007/008/009 using the model fit to Japanese subjects only. The packaged model contains the Japanese-fit parameters; it does not encode separate White-cohort estimates because none were estimated by the authors.
- Race recorded as Asian = 100%. Hayashi 2007 Table 1 enumerates 202 Japanese subjects in studies 1101 and 1305 with no other race groups in the model-building cohort; the population metadata records the cohort as 100% Asian (Japanese) accordingly.
-
Initial total_target uses observed IGE0, not the model
SS. As described in the Mechanism-check section, initialising
at IGE causes a small pre-treatment drift before binding dominates.
Users wanting a steady-state initialisation can override
inits = c(total_target = p_ige / cl_ige * v_ige)after computingp_ige,cl_ige, andv_igefor the cohort of interest.
Reproducibility
sessionInfo()
#> R version 4.6.0 (2026-04-24)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.4 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] PKNCA_0.12.1 ggplot2_4.0.3 tidyr_1.3.2
#> [4] dplyr_1.2.1 rxode2_5.1.2 nlmixr2lib_0.3.2.9000
#>
#> loaded via a namespace (and not attached):
#> [1] gtable_0.3.6 xfun_0.58 bslib_0.11.0
#> [4] lattice_0.22-9 vctrs_0.7.3 tools_4.6.0
#> [7] generics_0.1.4 parallel_4.6.0 tibble_3.3.1
#> [10] symengine_0.2.11 pkgconfig_2.0.3 data.table_1.18.4
#> [13] checkmate_2.3.4 RColorBrewer_1.1-3 S7_0.2.2
#> [16] desc_1.4.3 RcppParallel_5.1.11-2 lifecycle_1.0.5
#> [19] compiler_4.6.0 farver_2.1.2 textshaping_1.0.5
#> [22] fontawesome_0.5.3 htmltools_0.5.9 sys_3.4.3
#> [25] sass_0.4.10 yaml_2.3.12 crayon_1.5.3
#> [28] pillar_1.11.1 pkgdown_2.2.0 jquerylib_0.1.4
#> [31] whisker_0.4.1 openssl_2.4.1 cachem_1.1.0
#> [34] nlme_3.1-169 qs2_0.2.2 tidyselect_1.2.1
#> [37] digest_0.6.39 lotri_1.0.4 purrr_1.2.2
#> [40] labeling_0.4.3 rxode2ll_2.0.14 fastmap_1.2.0
#> [43] grid_4.6.0 cli_3.6.6 dparser_1.3.1-13
#> [46] magrittr_2.0.5 withr_3.0.2 scales_1.4.0
#> [49] backports_1.5.1 rmarkdown_2.31 otel_0.2.0
#> [52] askpass_1.2.1 ragg_1.5.2 stringfish_0.19.0
#> [55] memoise_2.0.1 evaluate_1.0.5 knitr_1.51
#> [58] rex_1.2.2 PreciseSums_0.7 rlang_1.2.0
#> [61] downlit_0.4.5 Rcpp_1.1.1-1.1 glue_1.8.1
#> [64] xml2_1.5.2 jsonlite_2.0.0 R6_2.6.1
#> [67] systemfonts_1.3.2 fs_2.1.0