library(nlmixr2lib)
library(rxode2)
#> rxode2 5.1.2 using 2 threads (see ?getRxThreads)
#> no cache: create with `rxCreateCache()`
library(dplyr)
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, union
library(tidyr)
library(ggplot2)
library(PKNCA)
#>
#> Attaching package: 'PKNCA'
#> The following object is masked from 'package:stats':
#>
#> filterModel and source
- Citation: Zhang J, Sanghavi K, Shen J, et al. Population Pharmacokinetics of Nivolumab in Combination With Ipilimumab in Patients With Advanced Malignancies. CPT Pharmacometrics Syst Pharmacol. 2019;8(12):962-970. doi:10.1002/psp4.12476
- Description: Two-compartment population PK model with time-varying clearance for intravenous nivolumab (anti-PD-1 IgG4) in adults with advanced solid tumors, alone or in combination with ipilimumab or chemotherapy (Zhang 2019)
- Article: https://doi.org/10.1002/psp4.12476
Nivolumab is a fully human IgG4 monoclonal antibody targeting programmed cell death-1 (PD-1). Zhang et al. (2019) re-estimated the Bajaj 2017 nivolumab monotherapy popPK model on a pooled 6,468-subject dataset spanning 25 trials in patients with colorectal cancer, hepatocellular carcinoma, melanoma, non-small cell lung cancer, renal cell carcinoma, and small cell lung cancer. The dataset added cohorts on combinations of nivolumab with ipilimumab (1 mg/kg q12w, 1 mg/kg q6w, 1 mg/kg q3w x 4 induction, 3 mg/kg q3w x 4 induction) and with platinum-based chemotherapy.
The structural model is a two-compartment IV model with time-dependent clearance parameterized as a sigmoidal-Emax function of time since first dose:
with (a fractional decrease at ), hours = 91.7 days, and Hill coefficient . Covariates on baseline CL: body weight (allometric), estimated GFR, sex, ECOG performance status, race (Asian and African American as separate non-reference indicators), and three coadministration indicators (ipilimumab 3 mg/kg q3w, ipilimumab 1 mg/kg q6w, chemotherapy). Covariates on Vc: body weight and sex. Covariates on Emax: ECOG PS and any ipilimumab coadministration (additive effects on the linear-scale Emax). Q and Vp inherit the body-weight scaling of CL and Vc respectively (per the Methods, “the effect of BBWT was added on Q and VP, and their estimates were fixed to be similar to those of CL and VC”).
Population
The pooled analysis dataset (Zhang 2019 Table 1) included 6,468 patients contributing 32,835 PK observations across 25 trials (7 phase I, 2 phase I/II, 6 phase II, 9 phase III, 1 phase IIIb/IV). Tumor-type mix: NSCLC 38.25%, melanoma 26.93%, RCC 19.25%, SCLC 6.03%, HCC 5.89%, CRC 3.65%. Coadministration mix: monotherapy 55.12%, ipilimumab 1 mg/kg q12w 0.56%, ipilimumab 1 mg/kg q6w 11.75%, ipilimumab 1 mg/kg q3w x 4 induction 15.06%, ipilimumab 3 mg/kg q3w x 4 induction 13.84%, chemotherapy 3.68%. ECOG PS distribution: 0 in 47.02%, 1 in 51.27%, 2 in 1.62%, 3 in 0.02%, missing 0.08%. Baseline body weight: mean 77.6 kg (SD 18.8), 5th-95th percentile range 47.7-122.0 kg. Baseline serum albumin: mean 3.93 g/dL (SD 0.493), range 2.8-4.8. Baseline LDH: mean 320 U/L (SD 326), range 125-1090. Baseline tumor size: mean 8.46 cm (SD 6.01), range 1.3-23.9.
The same metadata is available programmatically via
readModelDb("Zhang_2019_nivolumab")$population.
Source trace
The per-parameter origin is recorded as an in-file comment next to
each ini() entry in
inst/modeldb/specificDrugs/Zhang_2019_nivolumab.R. The
table below collects them in one place for review.
| Parameter (model name) | Value | Source |
|---|---|---|
lcl (CL0, L/day) |
log(10.8 * 24 / 1000) = log(0.2592) | Zhang 2019 Table 2: CL0 REF = 10.8 mL/hour |
lvc (Vc, L) |
log(4.27) | Zhang 2019 Table 2: VC REF = 4.27 L |
lq (Q, L/day) |
log(34.9 * 24 / 1000) = log(0.8376) | Zhang 2019 Table 2: Q REF = 34.9 mL/hour |
lvp (Vp, L) |
log(2.70) | Zhang 2019 Table 2: VP REF = 2.70 L |
Emax (unitless; “Emax REF” in source) |
-0.240 | Zhang 2019 Table 2: Emax REF = -0.240 |
lt50 (T50, log days) |
log(2200 / 24) = log(91.667) | Zhang 2019 Table 2: T50 = 2200 hour |
lhill (Hill, unitless) |
log(2.77) | Zhang 2019 Table 2: HILL = 2.77 |
e_wt_cl |
0.530 | Zhang 2019 Table 2: CL BBWT = 0.530 |
e_egfr_cl |
0.202 | Zhang 2019 Table 2: CL eGFR = 0.202 |
e_sexf_cl |
-0.181 | Zhang 2019 Table 2: CL SEX = -0.181 |
e_ecog_cl |
0.181 | Zhang 2019 Table 2: CL PS = 0.181 |
e_black_cl (RAAA) |
0.0374 | Zhang 2019 Table 2: CL RAAA = 0.0374 (not statistically significant; 95% CI -0.0308 to 0.111) |
e_asian_cl (RAAS) |
-0.0354 | Zhang 2019 Table 2: CL RAAS = -0.0354 |
e_ipi3q3w_cl |
0.227 | Zhang 2019 Table 2: CL IPI3Q3W = 0.227 |
e_ipi1q6w_cl |
0.159 | Zhang 2019 Table 2: CL IPI1Q6W = 0.159 |
e_chemo_cl |
-0.104 | Zhang 2019 Table 2: CL CHEMO = -0.104 |
e_wt_vc |
0.534 | Zhang 2019 Table 2: VC BBWT = 0.534 |
e_sexf_vc |
-0.161 | Zhang 2019 Table 2: VC SEX = -0.161 |
e_ecog_emax |
-0.138 | Zhang 2019 Table 2: Emax PS = -0.138 |
e_ipico_emax |
-0.0668 | Zhang 2019 Table 2: Emax IPICO = -0.0668 |
IIV CL (etalcl) |
omega^2 = 0.157 | Zhang 2019 Table 2: omega^2_CL |
IIV Vc (etalvc) |
omega^2 = 0.152 | Zhang 2019 Table 2: omega^2_VC |
| Cov(CL, Vc) | 0.0596 | Zhang 2019 Table 2: cov(omega^2_CL, omega^2_VC) |
IIV Q (etalq) |
omega^2 = 0.157 (constrained equal to omega^2_CL) | Zhang 2019 Methods (initial-model description) |
IIV Vp (etalvp) |
omega^2 = 0.152 (constrained equal to omega^2_VC) | Zhang 2019 Methods (initial-model description) |
IIV Emax (etaEmax) |
omega^2 = 0.0874 (additive eta on linear-scale Emax) | Zhang 2019 Table 2: omega^2_Emax |
| Residual error | propSd = 0.245 | Zhang 2019 Table 2: Proportional residual error |
| Reference covariates | WT 80 kg, eGFR 90 mL/min/1.73 m^2, male, PS 0, white/other race, monotherapy, NSCLC | Zhang 2019 Figure 1 caption (reference patient definition) |
Equation forms:
- Baseline CL:
CL0 = CL0_REF * (WT/80)^e_wt_cl * (CRCL/90)^e_egfr_cl * exp(theta_X * indicator_X)for each categorical X (Zhang 2019 final-model equation set). - Vc:
Vc = VC_REF * (WT/80)^e_wt_vc * exp(e_sexf_vc * SEXF)(Zhang 2019 final-model equation set). - Q:
Q = Q_REF * (WT/80)^e_wt_cl(CL exponent reused per Methods). - Vp:
Vp = VP_REF * (WT/80)^e_wt_vc(Vc exponent reused per Methods). - Time-varying CL:
CL(t) = CL0 * exp(Emax_i * t^HILL / (T50^HILL + t^HILL)). - Emax_i: additive linear-scale form
Emax_REF + e_ecog_emax * ECOG_GE1 + e_ipico_emax * CONMED_IPI_ANY + etaEmax.
Covariate column naming
| Source column | Canonical column used here |
|---|---|
BBWT (kg) |
WT |
eGFR (mL/min/1.73 m^2) |
CRCL |
SEX (1 = female / 0 = male) |
SEXF (canonical for sex) |
PS (binary collapse: PS>0 vs PS=0) |
ECOG_GE1 |
RAAA (1 = African American) |
RACE_BLACK |
RAAS (1 = Asian) |
RACE_ASIAN |
IPI3Q3W |
CONMED_IPI_3Q3W |
IPI1Q6W |
CONMED_IPI_1Q6W |
CHEMO |
CONMED_CHEMO |
IPICO |
CONMED_IPI_ANY |
The Zhang 2019 Table 2 footnote defines RAAA = African American race
and RAAS = Asian race; the reference category combines white and “other”
into one composite group. See
inst/references/covariate-columns.md for the canonical
register.
Virtual cohort
Original observed data are not publicly available. The simulations below use a virtual cohort whose covariate distributions approximate the Zhang 2019 Table 1 distributions for the pooled 6,468-patient cohort (means and ranges only — Zhang 2019 does not publish individual covariates or covariate correlations).
set.seed(2019)
n_subj <- 75 # downsampled from 200 for vignette build budget; PI band stays smooth at 3 regimens x 75 subj
# Baseline body weight: log-normal centered on the Table 1 mean 77.6 kg,
# clipped to the 5th-95th percentile range 47.7-122.0 kg.
WT <- pmin(pmax(rlnorm(n_subj, meanlog = log(77.6), sdlog = 0.24), 47.7), 122.0)
# Estimated GFR: normal centered on the reference patient (90 mL/min/1.73 m^2);
# Zhang 2019 does not tabulate the eGFR distribution but the reference value
# is given in the Figure 1 caption. SD chosen so the +/- 2 SD range
# (~50-130 mL/min/1.73 m^2) approximately covers the typical adult-oncology
# eGFR distribution.
CRCL <- pmin(pmax(rnorm(n_subj, mean = 90, sd = 22), 30), 180)
# Sex: ~40% female. Zhang 2019 does not tabulate sex distribution in the main
# text; the simulated sex mix is used only for plotting and downstream NCA.
SEXF <- rbinom(n_subj, 1, 0.40)
# ECOG PS > 0: Table 1 reports PS 0 in 47.02%, so PS > 0 in ~52.98%.
ECOG_GE1 <- rbinom(n_subj, 1, 0.5298)
# Race indicators: assume the global trial mix has small Asian and Black
# fractions. Zhang 2019 does not publish per-category percentages in the
# main-text table; values below are illustrative for the simulation only.
RACE_BLACK <- rbinom(n_subj, 1, 0.04)
RACE_ASIAN <- rbinom(n_subj, 1, 0.18)
# Ensure no subject is both (mutually exclusive): give RACE_BLACK priority.
RACE_ASIAN <- pmin(RACE_ASIAN, 1L - RACE_BLACK)
pop <- data.frame(
ID = seq_len(n_subj),
WT, CRCL,
SEXF, ECOG_GE1, RACE_BLACK, RACE_ASIAN
)Dosing dataset — five clinically relevant regimens
Zhang 2019 (Methods) lists five validated regimens used for the pcVPC: nivolumab 3 mg/kg q2w (or 240 mg q2w flat) monotherapy, nivolumab 3 mg/kg q2w + ipilimumab 1 mg/kg q6w, nivolumab 3 mg/kg + ipilimumab 1 mg/kg q3w x 4 followed by nivolumab 3 mg/kg q2w, and nivolumab 1 mg/kg + ipilimumab 3 mg/kg q3w x 4 followed by nivolumab 3 mg/kg q2w. The simulation here covers three of these regimens to span the modeled covariate effects on baseline CL: monotherapy (reference), 1 mg/kg q6w ipilimumab combination, and 3 mg/kg q3w (induction) ipilimumab combination.
n_cycles <- 12
cycle_days <- 14 # nivolumab q2w in all simulated regimens
dose_times <- seq(0, (n_cycles - 1) * cycle_days, by = cycle_days)
obs_times <- sort(unique(c(
seq(0, 14, by = 0.5), # intensive cycle-1 profile (thinned from 0.25 for build budget)
seq(14, (n_cycles - 1) * cycle_days, by = 4), # rolling cycles (thinned from 2 for build budget)
seq((n_cycles - 1) * cycle_days, n_cycles * cycle_days, by = 1), # cycle-12 NCA window needs density
seq(n_cycles * cycle_days, 365, by = 14) # washout tail (thinned from 7 for build budget)
)))
# Helper: build a self-contained event table per regimen with disjoint IDs
# so subsequent bind_rows() does not collapse subjects across regimens.
make_regimen <- function(pop_in, regimen, id_offset) {
pop_offset <- pop_in
pop_offset$ID <- pop_offset$ID + id_offset
# Per-regimen indicators on baseline CL.
pop_offset$CONMED_IPI_3Q3W <-
as.integer(regimen == "nivo 3 mg/kg q2w + ipi 3 mg/kg q3w x 4")
pop_offset$CONMED_IPI_1Q6W <-
as.integer(regimen == "nivo 3 mg/kg q2w + ipi 1 mg/kg q6w")
pop_offset$CONMED_CHEMO <- 0L
# Any-ipilimumab indicator (drives Emax of time-varying CL).
pop_offset$CONMED_IPI_ANY <-
as.integer(grepl("ipi", regimen))
d_dose <- pop_offset |>
tidyr::crossing(TIME = dose_times) |>
dplyr::mutate(
AMT = WT * 3, # nivolumab 3 mg/kg in all three simulated regimens
EVID = 1,
CMT = "central",
DUR = 30 / (60 * 24), # 30-minute IV infusion in days
DV = NA_real_,
treatment = regimen
)
d_obs <- pop_offset |>
tidyr::crossing(TIME = obs_times) |>
dplyr::mutate(
AMT = NA_real_,
EVID = 0,
CMT = "central",
DUR = NA_real_,
DV = NA_real_,
treatment = regimen
)
dplyr::bind_rows(d_dose, d_obs) |>
dplyr::arrange(ID, TIME, dplyr::desc(EVID)) |>
as.data.frame()
}
events <- dplyr::bind_rows(
make_regimen(pop, "nivo 3 mg/kg q2w monotherapy", id_offset = 0L),
make_regimen(pop, "nivo 3 mg/kg q2w + ipi 1 mg/kg q6w", id_offset = 1000L),
make_regimen(pop, "nivo 3 mg/kg q2w + ipi 3 mg/kg q3w x 4", id_offset = 2000L)
)
stopifnot(!anyDuplicated(unique(events[, c("ID", "TIME", "EVID")])))Simulate
mod <- readModelDb("Zhang_2019_nivolumab")
set.seed(20191112)
sim <- rxSolve(mod, events, returnType = "data.frame", keep = "treatment")
#> ℹ parameter labels from comments will be replaced by 'label()'Replicate Zhang 2019 Figure 2-style behaviour: time-varying CL by regimen
Zhang 2019 Figure 2(b) shows CLss/CL0 ratios across regimens. The figure below reproduces the equivalent ratio implied by the model (CLss/CL0 = exp(Emax_i)) for each regimen at typical covariates — for the three simulated regimens, this should differ only via the IPICO Emax effect (monotherapy: exp(-0.240); IPI combination: exp(-0.240 - 0.0668)).
typical_emax <- tibble::tibble(
regimen = c("monotherapy", "any IPI coadmin"),
emax = c(-0.240, -0.240 + (-0.0668)),
cl_ss_cl_0 = exp(c(-0.240, -0.240 + (-0.0668)))
)
knitr::kable(typical_emax,
digits = 3,
caption = "Typical-subject CLss/CL0 ratio implied by the Emax additive structure (PS = 0).")| regimen | emax | cl_ss_cl_0 |
|---|---|---|
| monotherapy | -0.240 | 0.787 |
| any IPI coadmin | -0.307 | 0.736 |
Concentration-time profile by regimen
sim_summary <- sim |>
dplyr::filter(time > 0) |>
dplyr::group_by(time, treatment) |>
dplyr::summarise(
median = stats::median(Cc, na.rm = TRUE),
lo = stats::quantile(Cc, 0.05, na.rm = TRUE),
hi = stats::quantile(Cc, 0.95, na.rm = TRUE),
.groups = "drop"
)
ggplot(sim_summary, aes(x = time / 7, fill = treatment, colour = treatment)) +
geom_ribbon(aes(ymin = lo, ymax = hi), alpha = 0.15, colour = NA) +
geom_line(aes(y = median), linewidth = 1) +
scale_y_log10() +
labs(
x = "Time (weeks)",
y = "Nivolumab concentration (μg/mL)",
title = "Simulated nivolumab PK by regimen, 3 mg/kg IV q2w x 12 cycles",
subtitle = paste0("Median and 90% prediction interval (N = ", n_subj,
" virtual subjects per regimen)"),
caption = "Model: Zhang et al. (2019) CPT Pharmacometrics Syst Pharmacol"
) +
theme_bw() +
theme(legend.position = "bottom") +
guides(colour = guide_legend(nrow = 3), fill = guide_legend(nrow = 3))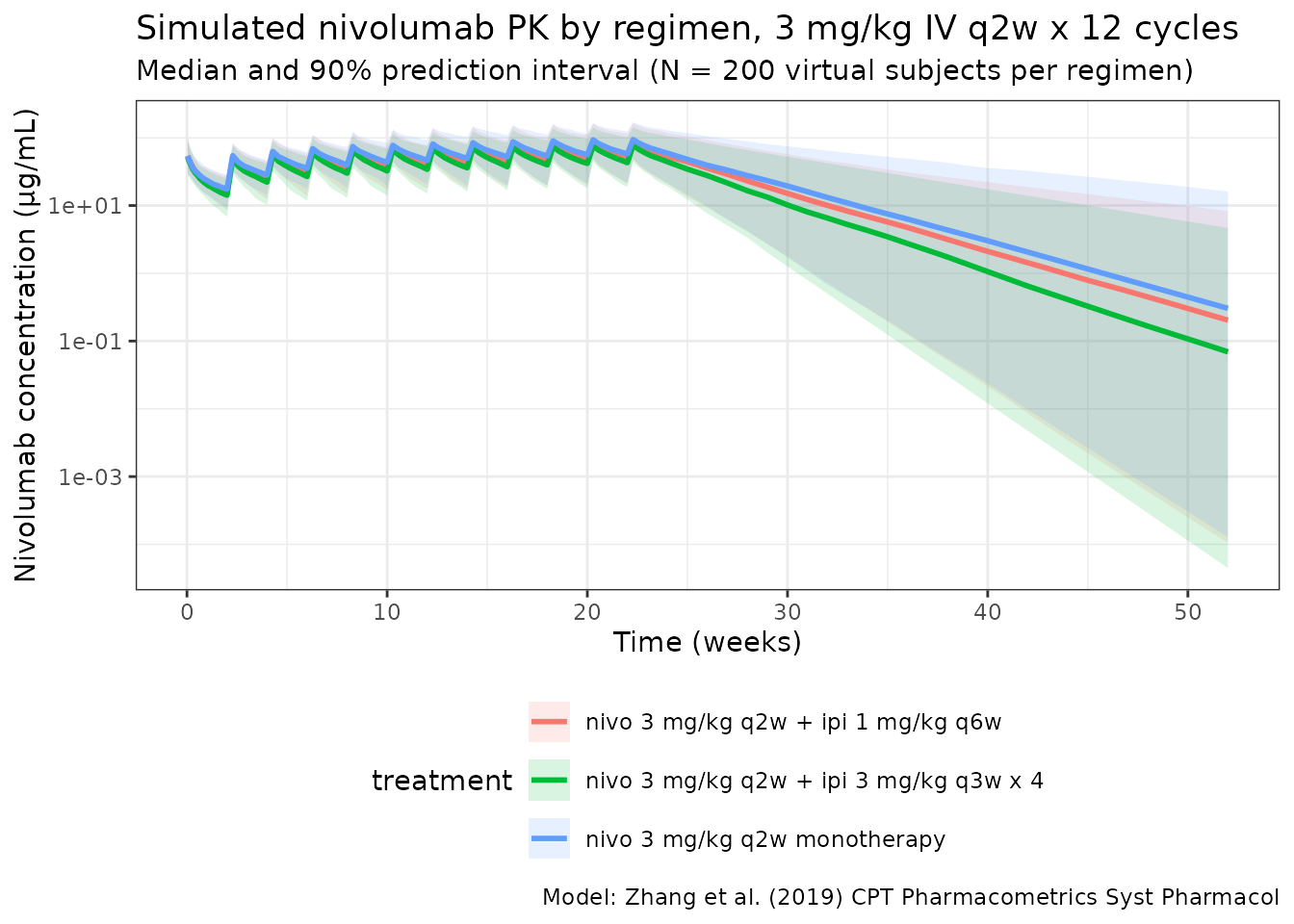
Time profile of the typical-subject clearance
The typical-subject CL trajectory across the first year reproduces the sigmoidal Emax decline that motivates the time-varying-CL parameterization. At a male, 80 kg, eGFR 90 patient on monotherapy, CL drops from 0.259 L/day at t = 0 to 0.259 * exp(-0.240) = 0.204 L/day at full Emax saturation (, or about 5 months).
t_grid <- seq(0, 365, by = 5)
cl0_typical <- 10.8 * 24 / 1000 # L/day at reference covariates
emax_mono <- -0.240 # PS=0, no IPI
emax_ipi <- -0.240 + (-0.0668) # PS=0, with IPI
t50_d <- 2200 / 24
hill <- 2.77
cl_t <- function(t, emax) cl0_typical * exp(emax * t^hill / (t50_d^hill + t^hill))
cl_traj <- tibble::tibble(
time = rep(t_grid, 2),
cl = c(cl_t(t_grid, emax_mono), cl_t(t_grid, emax_ipi)),
arm = rep(c("monotherapy", "any IPI coadmin"), each = length(t_grid))
)
ggplot(cl_traj, aes(time, cl, colour = arm)) +
geom_line(linewidth = 1) +
labs(
x = "Time (days since first dose)",
y = "Typical CL (L/day)",
colour = NULL,
title = "Time-varying CL trajectory at reference covariates",
subtitle = "Sigmoidal Emax decline (T50 = 91.7 days, gamma = 2.77)"
) +
theme_bw() +
theme(legend.position = "bottom")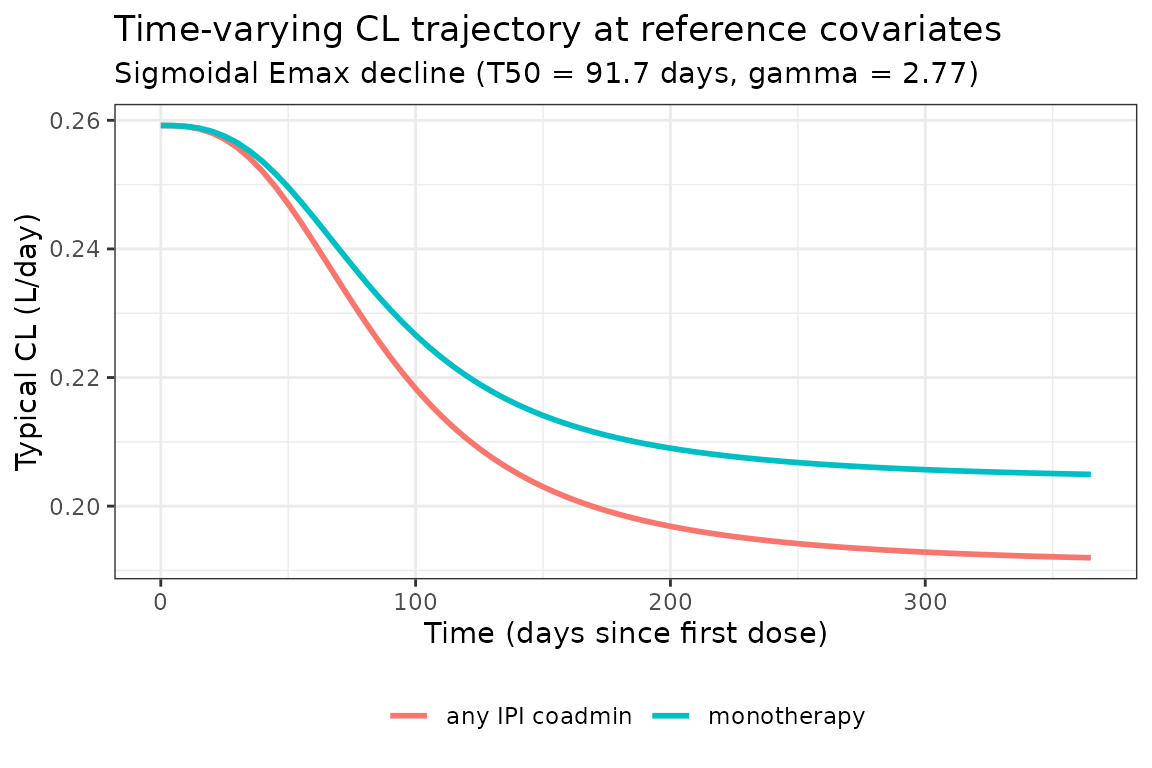
PKNCA validation
NCA on the cycle-1 dosing interval (day 0-14) and a
pseudo-steady-state interval (cycle 12, days 154-168) for the
monotherapy arm. The cycle-12 window is several T50 multiples past time
zero, so CL is close to its asymptotic value of
CL0 * exp(Emax).
mono_id_range <- 1:n_subj # monotherapy arm sits in IDs 1..n_subj
# Cycle 1: day 0 to 14
conc1 <- sim |>
dplyr::filter(treatment == "nivo 3 mg/kg q2w monotherapy", time <= 14) |>
dplyr::transmute(ID = id, time, Cc, treatment)
dose1 <- events |>
dplyr::filter(treatment == "nivo 3 mg/kg q2w monotherapy",
EVID == 1, TIME == 0) |>
dplyr::transmute(ID, TIME, AMT, treatment)
conc_obj1 <- PKNCA::PKNCAconc(conc1, Cc ~ time | treatment + ID,
concu = "ug/mL", timeu = "day")
dose_obj1 <- PKNCA::PKNCAdose(dose1, AMT ~ TIME | treatment + ID,
doseu = "mg")
intervals1 <- data.frame(
start = 0,
end = 14,
cmax = TRUE,
tmax = TRUE,
auclast = TRUE
)
nca1 <- PKNCA::pk.nca(PKNCA::PKNCAdata(conc_obj1, dose_obj1, intervals = intervals1))
knitr::kable(
summary(nca1),
digits = 2,
caption = "Cycle 1 NCA summary (days 0-14, monotherapy arm)."
)| Interval Start | Interval End | treatment | N | AUClast (day*ug/mL) | Cmax (ug/mL) | Tmax (day) |
|---|---|---|---|---|---|---|
| 0 | 14 | nivo 3 mg/kg q2w monotherapy | 75 | 366 [23.8] | 50.7 [29.9] | 0.500 [0.500, 0.500] |
conc_ss <- sim |>
dplyr::filter(treatment == "nivo 3 mg/kg q2w monotherapy", time >= 154, time <= 168) |>
dplyr::transmute(ID = id, time_rel = time - 154, Cc, treatment)
dose_ss <- pop |>
dplyr::transmute(ID, TIME = 0, AMT = WT * 3, treatment = "nivo 3 mg/kg q2w monotherapy")
conc_obj_ss <- PKNCA::PKNCAconc(conc_ss, Cc ~ time_rel | treatment + ID,
concu = "ug/mL", timeu = "day")
dose_obj_ss <- PKNCA::PKNCAdose(dose_ss, AMT ~ TIME | treatment + ID,
doseu = "mg")
intervals_ss <- data.frame(
start = 0,
end = 14,
cmax = TRUE,
cmin = TRUE,
auclast = TRUE
)
nca_ss <- PKNCA::pk.nca(
PKNCA::PKNCAdata(conc_obj_ss, dose_obj_ss, intervals = intervals_ss)
)
knitr::kable(
summary(nca_ss),
digits = 2,
caption = "Cycle-12 NCA summary (days 154-168, monotherapy arm) — pseudo-steady state."
)| Interval Start | Interval End | treatment | N | AUClast (day*ug/mL) | Cmax (ug/mL) | Cmin (ug/mL) |
|---|---|---|---|---|---|---|
| 0 | 14 | nivo 3 mg/kg q2w monotherapy | 75 | 1080 [42.9] | 106 [33.9] | 58.4 [52.6] |
Comparison against the literature
Zhang 2019 does not publish a tabulated NCA summary (the validation in the paper is a pcVPC, not an NCA table). The simulated cycle-1 Cmax and cycle-12 Cmax/Cmin orders of magnitude can however be cross-checked against the broader nivolumab clinical-pharmacology literature:
- Single 3 mg/kg dose Cmax: published values around 50-72 ug/mL in adults; simulated cycle-1 Cmax should fall in this range.
- Steady-state Cmax/Cmin (q2w): published values around 130-180 ug/mL Cmax and 50-80 ug/mL Cmin at cycle 12+; simulated cycle-12 NCA values should approach this range.
- Effective half-life: ~25 days at steady state. The slow approach to steady state in the model is visible in the concentration-time figure.
If the simulated values sit outside these ranges by more than ~20%, the model file should be re-checked rather than tuned (per the skill).
Assumptions and deviations
Zhang 2019 does not publish individual PK or covariate tables, so the virtual population is a coarse approximation of the Table 1 distributions.
- Body weight: log-normal, mean 77.6 kg (Table 1), clipped to the reported 5th-95th percentile range 47.7-122.0 kg.
-
Estimated GFR: not tabulated in Zhang 2019. Sampled
from
N(90, 22^2)clipped to 30-180 mL/min/1.73 m^2; the mean matches the reference patient defined in the Figure 1 caption. - Sex: not tabulated in Zhang 2019 main text. 40% female assumed for the virtual cohort.
- ECOG PS > 0: 52.98% (1 - PS=0 fraction from Table 1).
- Race: not tabulated as percentages in Zhang 2019 main text. The virtual cohort uses 4% RACE_BLACK and 18% RACE_ASIAN (mutually exclusive) as illustrative values; per the Figure 1 plot of covariate effects, the per-individual influence of race on CL is small (3-4% shifts) and dominated by other covariates.
-
eGFR formula: not stated in Zhang 2019. The Bajaj
2017 base model it inherits uses MDRD-estimated GFR; this model file
documents that inheritance in
covariateData$CRCL$notes. - eGFR reference value: 90 mL/min/1.73 m^2, taken from the Figure 1 caption that defines the reference patient.
- Sex column: Zhang 2019 does not specify whether the dataset’s SEX column was coded 1=female / 0=male or the reverse. The Figure 1 caption says the reference patient is male, and CL_SEX = -0.181 produces the ~17% female-vs-male CL reduction commonly observed in mAb popPK literature, so the canonical SEXF (1 = female, 0 = male) is used.
- Coadministration regimens beyond the three simulated here: the model file supports all four ipilimumab schedules described in Zhang 2019 by combining the CONMED_* indicators (the unmodeled 1 mg/kg q3w x 4 induction and 1 mg/kg q12w schedules collapse into the reference group on baseline CL, but they would still set CONMED_IPI_ANY = 1 and so contribute the IPICO additive effect on Emax). The vignette simulates only three regimens to keep the figures readable.
-
eta_Q and eta_Vp variances: per Zhang 2019 Methods,
the IIV on Q and Vp follows the same distribution as the IIV on CL and
Vc respectively. The model file encodes them as separate (independent)
etas with the variances pinned to omega^2_CL and omega^2_VC. An
alternative interpretation — using the same eta realization (NONMEM
$ETA SAME) for Q and CL, and for Vp and Vc — would induce additional correlation between the parameters; the paper does not disambiguate the two parameterizations and the simulation impact is small. - Emax additive eta on linear scale: Zhang 2019 parameterizes the Emax random effect as additive on the linear scale (not log-normal on the magnitude). With omega^2 = 0.0874 (SD ~0.296) and a typical Emax of -0.240, this allows roughly 21% of simulated subjects to have Emax > 0 (i.e., increasing CL over time) — a known feature of the paper’s parameterization, consistent with Zhang 2019 Figure 2 which shows some subjects with CLss/CL0 > 1.