Ipilimumab (Sanghavi 2020)
Source:vignettes/articles/Sanghavi_2020_ipilimumab.Rmd
Sanghavi_2020_ipilimumab.RmdModel and source
- Citation: Sanghavi K, Zhang J, Zhao X, et al. Population Pharmacokinetics of Ipilimumab in Combination With Nivolumab in Patients With Advanced Solid Tumors. CPT Pharmacometrics Syst Pharmacol. 2020;9(1):29-39. doi:10.1002/psp4.12477
- Description: Two-compartment population PK model for intravenous ipilimumab (anti-CTLA-4 IgG1) with time-varying clearance via a sigmoid emax function in patients with advanced solid tumors receiving ipilimumab alone or in combination with nivolumab (Sanghavi 2020)
- Article: https://doi.org/10.1002/psp4.12477
Ipilimumab is a fully human anti-CTLA-4 IgG1 monoclonal antibody. The Sanghavi 2020 analysis refines an earlier ipilimumab population PK model (Feng 2014, reference 17 in the paper) with a much larger combination-therapy dataset and a sigmoid-Emax description of time-varying clearance.
Structural form: linear two-compartment IV model with first-order elimination and a multiplicative time-on-CL term:
where is negative for a typical patient on combination therapy (CL decreases over time), days, and gives a sharp sigmoid switch between baseline and steady-state CL. Baseline body weight (BBWT) scales CL and Q as and VC and VP as .
Population
The final-model population pooled 3,411 patients with advanced solid tumors across 16 clinical trials (12,545 ipilimumab serum concentrations). The breakdown (Sanghavi 2020 Table 1):
- Tumor type: melanoma 50.4%, non-small cell lung cancer 17.2%, renal cell carcinoma 13.1%, small cell lung cancer 5.2%, hepatocellular carcinoma 3.8%, colorectal cancer 3.6%.
- Treatment: ipilimumab monotherapy 26.2% (N = 893); ipilimumab in combination with nivolumab 73.8% (N = 2,518) across five nivolumab regimens (0.3 mg/kg Q3W, 1 mg/kg Q2W, 1 mg/kg Q3W, 3 mg/kg Q2W, 3 mg/kg Q3W).
- Demographics: body weight median 76.8 kg (range 36.8–181); baseline LDH median 217 U/L (range 74–6,245); baseline albumin median 4.1 g/dL (range 1.8–5.3); baseline tumor size median 6.29 cm. Performance status 0 in 57.3%, 1 in 41.3%, ≥ 2 in 1.4%.
- Trials: two phase I, two phase I/II, eight phase II, three phase III, one phase IIIb/IV (multinational).
The same metadata is available programmatically via
rxode2::rxode(readModelDb("Sanghavi_2020_ipilimumab"))$population.
Source trace
The per-parameter origin is recorded as an in-file comment next to
each ini() entry in
inst/modeldb/specificDrugs/Sanghavi_2020_ipilimumab.R. The
table below collects them in one place for review.
| Parameter (model name) | Value (this package) | Source location |
|---|---|---|
lcl (CL0, L/day) |
log(14.1 × 0.024) | Table 2: CL0_REF = 14.1 mL/h |
lvc (VC, L) |
log(3.95) | Table 2: VC_REF |
lq (Q, L/day) |
log(27.9 × 0.024) | Table 2: Q_REF = 27.9 mL/h |
lvp (VP, L) |
log(3.18) | Table 2: VP_REF |
Emax (monotherapy) |
-0.0644 | Table 2: Emax_REF |
lt50 (T50, days) |
log(2540 / 24) = log(105.83) | Table 2: T50 = 2,540 h |
lhill (HILL exponent) |
log(7.43) | Table 2: HILL |
e_wt_cl_q (WT on CL/Q) |
0.694 | Table 2: CL_BBWT |
e_wt_vc_vp (WT on Vc/Vp) |
0.600 | Table 2: V_BBWT |
e_logldh_cl |
0.703 | Table 2: CL_log-BLDH |
e_sclc_cl |
-0.124 | Table 2: CL_SCLC |
e_line_cl (1L vs 2L+) |
-0.0949 | Table 2: CL_LINE |
e_n1q3w_cl |
0.0950 | Table 2: CL_N1Q3W |
e_n3q2w_cl |
0.191 | Table 2: CL_N3Q2W |
e_combo_emax |
-0.202 | Table 2: Emax_COMBO |
IIV etalcl + etalvc block |
c(0.112, 0.0404, 0.0884) | Table 2: ω²_CL, cov, ω²_VC |
etaEmax |
0.0158 | Table 2: ω²_Emax |
propSd |
0.223 | Table 2: Proportional |
addSd (μg/mL) |
0.607 | Table 2: Additive |
Equations (from the paper’s “Final model” subsection in Results):
cl0 = CL0_REF · (BBWT/80)^CL_BBWT · (log(BLDH)/log(217))^CL_logBLDH · exp(CL_SCLC·I_SCLC) · exp(CL_LINE·I_1L) · exp(CL_N1Q3W·I_N1Q3W) · exp(CL_N3Q2W·I_N3Q2W) · exp(η_CL)Emax_i = Emax_REF + Emax_COMBO·I_COMBO + η_Emaxcl(t) = cl0_i · exp(Emax_i · t^HILL / (T50^HILL + t^HILL))vc = VC_REF · (BBWT/80)^V_BBWT · exp(η_VC)q = Q_REF · (BBWT/80)^CL_BBWTvp = VP_REF · (BBWT/80)^V_BBWT
Virtual cohort
Original observed data are not publicly available. The simulations below use a virtual melanoma cohort whose body-weight distribution approximates the pooled trial population (median 76.8 kg, range 36.8–181 kg), with reference covariates fixed at the paper’s reference patient values (LDH 217 U/L, melanoma, 2L+) so the time-on-CL component is the dominant driver of between-cohort differences.
set.seed(2020)
n_subj <- 100 # downsampled from 200 for vignette build budget; 90% PI band shape preserved
cohort <- tibble(
ID = seq_len(n_subj),
WT = pmin(pmax(rlnorm(n_subj, log(76.8), 0.27), 36.8), 181),
LDH = 217,
TUMTP_SCLC = 0L, # melanoma reference (SCLC indicator off)
LINE_1L = 0L # 2L+ reference
)The Model Application section of Sanghavi 2020 simulates the approved combination induction: ipilimumab 3 mg/kg Q3W plus nivolumab 1 mg/kg Q3W, four doses. Two arms are compared here:
-
Combination: ipilimumab 3 mg/kg + nivolumab 1 mg/kg
Q3W × 4 (
COMBO_NIVO = 1,NIVO_1Q3W = 1). -
Monotherapy: ipilimumab 3 mg/kg Q3W × 4 alone
(
COMBO_NIVO = 0,NIVO_1Q3W = 0).
make_arm <- function(pop, regimen, id_offset = 0L) {
combo <- as.integer(regimen == "Combination")
n1q3w <- combo
ipi_amt <- pop$WT * 3 # 3 mg/kg
dose_t <- seq(0, by = 21, length.out = 4)
obs_t <- sort(unique(c(seq(0, 168, by = 1)))) # daily; 168 pts vs 672 (every 6 h)
d_dose <- pop |>
mutate(ID = id_offset + ID) |>
crossing(TIME = dose_t) |>
mutate(AMT = rep(ipi_amt, length(dose_t)),
EVID = 1, CMT = "central", DUR = 1.5 / 24, DV = NA_real_,
treatment = regimen,
NIVO_1Q3W = n1q3w, NIVO_3Q2W = 0L, COMBO_NIVO = combo)
d_obs <- pop |>
mutate(ID = id_offset + ID) |>
crossing(TIME = obs_t) |>
mutate(AMT = NA_real_, EVID = 0, CMT = "central", DUR = NA_real_,
DV = NA_real_, treatment = regimen,
NIVO_1Q3W = n1q3w, NIVO_3Q2W = 0L, COMBO_NIVO = combo)
bind_rows(d_dose, d_obs) |> arrange(ID, TIME, desc(EVID)) |> as.data.frame()
}
events <- bind_rows(
make_arm(cohort, "Combination", id_offset = 0L),
make_arm(cohort, "Monotherapy", id_offset = n_subj)
)
stopifnot(!anyDuplicated(unique(events[, c("ID", "TIME", "EVID")])))Simulation
mod <- readModelDb("Sanghavi_2020_ipilimumab")
sim <- rxode2::rxSolve(mod, events = events, returnType = "data.frame",
keep = c("treatment", "WT"))
#> ℹ parameter labels from comments will be replaced by 'label()'For deterministic typical-value replication the model’s random effects can be zeroed:
mod_typical <- mod |> rxode2::zeroRe()
#> ℹ parameter labels from comments will be replaced by 'label()'
events_ref <- data.frame(
ID = 1L, TIME = c(seq(0, 168, by = 0.5)),
AMT = NA_real_, EVID = 0L, CMT = "central", DUR = NA_real_,
DV = NA_real_, WT = 80, LDH = 217, TUMTP_SCLC = 0L, LINE_1L = 0L,
NIVO_1Q3W = 1L, NIVO_3Q2W = 0L, COMBO_NIVO = 1L
)
events_ref <- bind_rows(
data.frame(ID = 1L, TIME = seq(0, by = 21, length.out = 4),
AMT = 240, EVID = 1L, CMT = "central", DUR = 1.5 / 24,
DV = NA_real_, WT = 80, LDH = 217,
TUMTP_SCLC = 0L, LINE_1L = 0L,
NIVO_1Q3W = 1L, NIVO_3Q2W = 0L, COMBO_NIVO = 1L),
events_ref
) |> arrange(TIME, desc(EVID))
sim_ref <- rxode2::rxSolve(mod_typical, events = events_ref,
returnType = "data.frame")
#> ℹ omega/sigma items treated as zero: 'etalcl', 'etalvc', 'etaemax'Replicate published figures
Time-varying clearance (Figure 3, left side: monotherapy vs. combination)
Sanghavi 2020 Figure 3 reports the model-estimated CL trajectories by treatment arm. The plot below shows the typical-individual CL ratio for the two reference scenarios (80 kg melanoma 2L+ patient, LDH 217 U/L, no nivolumab regimen indicator on baseline CL):
t_grid <- seq(0, 365, by = 5)
make_typical_arm <- function(combo) {
ev <- data.frame(
ID = 1L,
TIME = c(seq(0, by = 21, length.out = 4), t_grid),
AMT = c(rep(240, 4), rep(NA_real_, length(t_grid))),
EVID = c(rep(1L, 4), rep(0L, length(t_grid))),
CMT = "central",
DUR = c(rep(1.5 / 24, 4), rep(NA_real_, length(t_grid))),
DV = NA_real_,
WT = 80, LDH = 217, TUMTP_SCLC = 0L, LINE_1L = 0L,
NIVO_1Q3W = combo, NIVO_3Q2W = 0L, COMBO_NIVO = combo
) |> arrange(TIME, desc(EVID))
res <- rxode2::rxSolve(mod_typical, events = ev, returnType = "data.frame")
res$treatment <- if (combo == 1) "Combination (nivo 1 mg/kg Q3W)" else "Monotherapy"
res[res$time > 0, ]
}
cl_traj <- bind_rows(make_typical_arm(0L), make_typical_arm(1L))
#> ℹ omega/sigma items treated as zero: 'etalcl', 'etalvc', 'etaemax'
#> ℹ omega/sigma items treated as zero: 'etalcl', 'etalvc', 'etaemax'
ggplot(cl_traj, aes(time, cl / cl0, colour = treatment)) +
geom_line(linewidth = 1) +
geom_hline(yintercept = 1, linetype = "dashed", colour = "grey40") +
labs(x = "Time since first dose (days)",
y = "CL(t) / CL(0)",
title = "Time-varying clearance trajectory (typical 80 kg patient)",
subtitle = "Replicates the qualitative pattern of Figure 3 of Sanghavi 2020",
colour = NULL) +
theme_bw() + theme(legend.position = "top")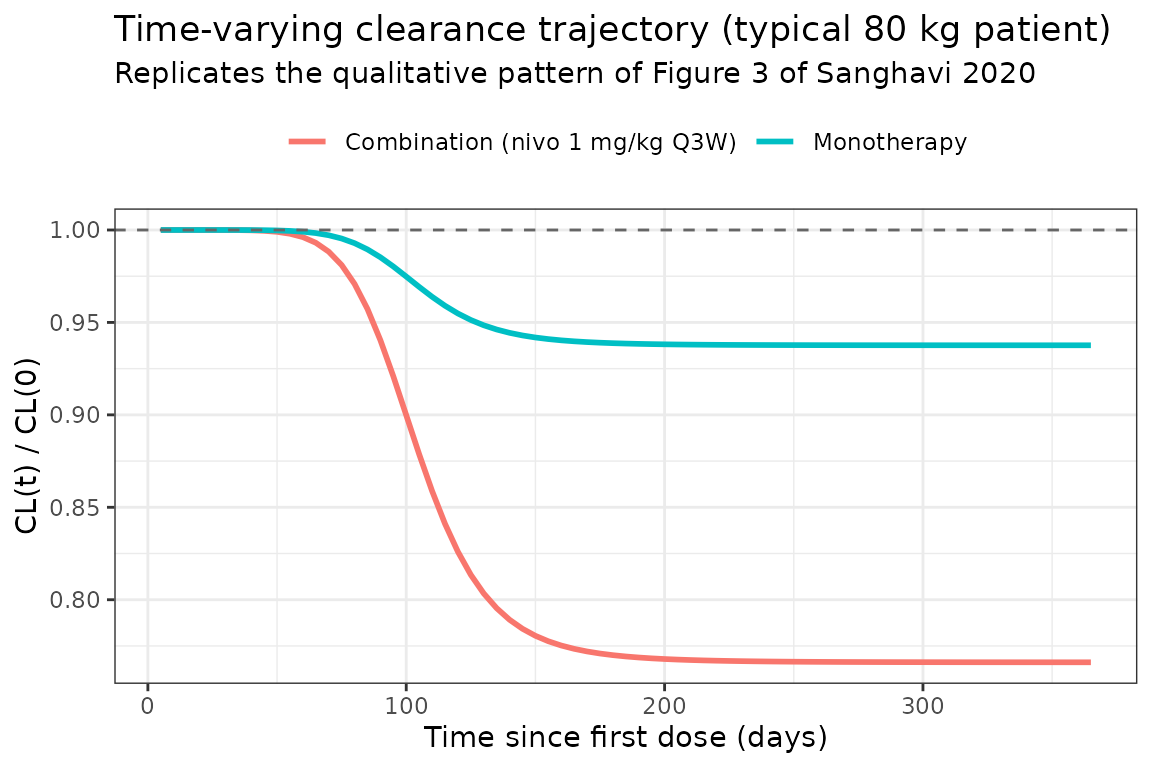
Combination-arm concentration profile
sim_summary <- sim |>
filter(time > 0) |>
group_by(time, treatment) |>
summarise(median = median(Cc, na.rm = TRUE),
lo = quantile(Cc, 0.05, na.rm = TRUE),
hi = quantile(Cc, 0.95, na.rm = TRUE),
.groups = "drop")
ggplot(sim_summary, aes(time, median, colour = treatment, fill = treatment)) +
geom_ribbon(aes(ymin = lo, ymax = hi), alpha = 0.15, colour = NA) +
geom_line(linewidth = 0.8) +
labs(x = "Time (days)", y = "Ipilimumab Cc (μg/mL)",
title = "Simulated ipilimumab PK: 3 mg/kg Q3W induction × 4 doses",
subtitle = paste0("Median and 90% prediction interval, N = ",
n_subj, " virtual subjects per arm"),
caption = "Model: Sanghavi 2020 CPT:PSP 9(1):29-39") +
theme_bw()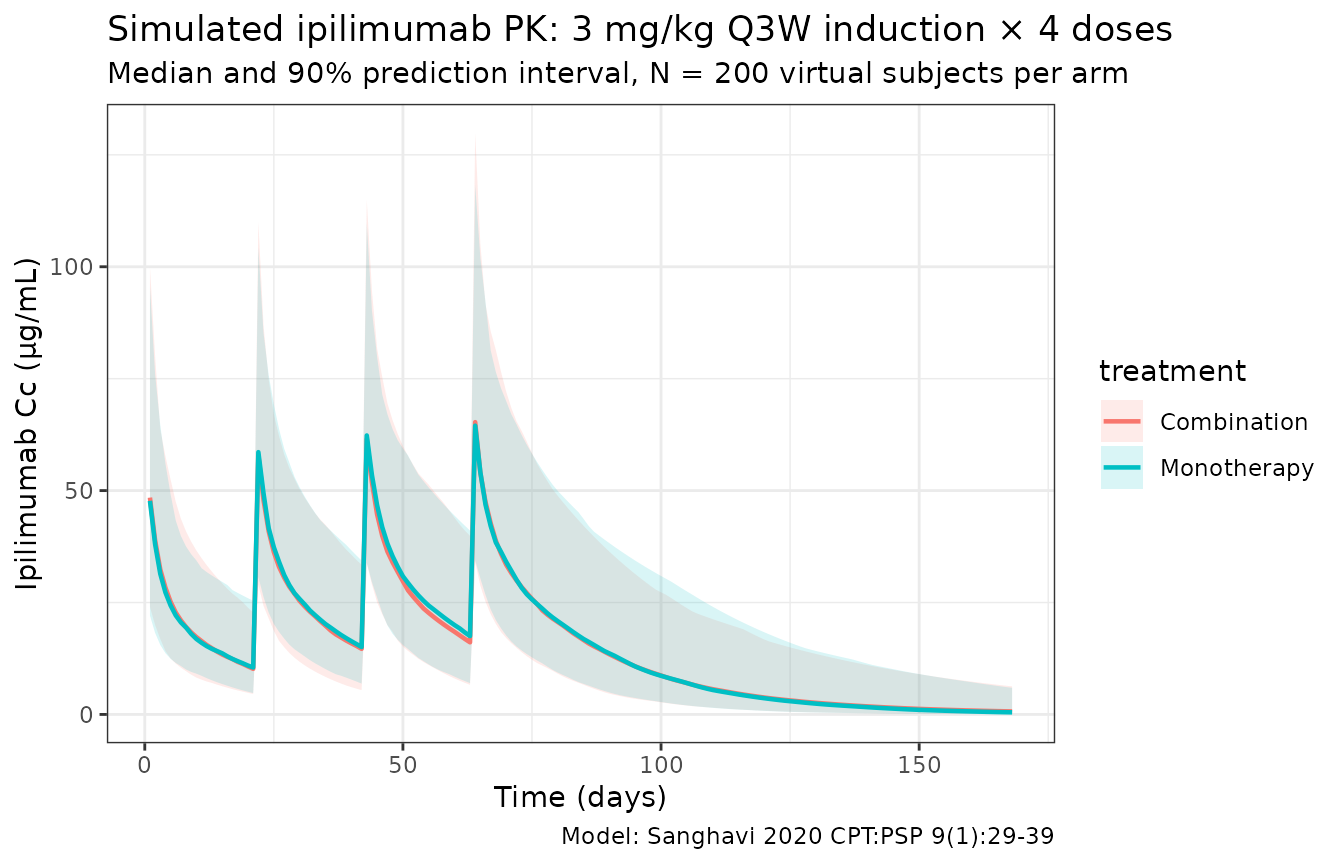
PKNCA validation
Compute NCA parameters over the first dosing interval (Day 0 to Day 21) for the combination arm, where Sanghavi 2020 Table 4 reports the geometric-mean Day-1 peak (62.6 μg/mL) and Day-21 trough (10.4 μg/mL).
sim_combo <- sim |>
filter(treatment == "Combination", !is.na(Cc), time <= 21) |>
select(id, time, Cc, treatment)
conc_obj <- PKNCA::PKNCAconc(sim_combo, Cc ~ time | treatment + id)
dose_df <- events |>
filter(EVID == 1, treatment == "Combination", TIME == 0) |>
transmute(id = ID, time = TIME, amt = AMT, treatment = treatment)
dose_obj <- PKNCA::PKNCAdose(dose_df, amt ~ time | treatment + id)
intervals <- data.frame(
start = 0, end = 21,
cmax = TRUE, tmax = TRUE,
auclast = TRUE, half.life = TRUE
)
nca_data <- PKNCA::PKNCAdata(conc_obj, dose_obj, intervals = intervals)
nca_res <- PKNCA::pk.nca(nca_data)
knitr::kable(summary(nca_res),
caption = "Simulated NCA parameters (combination arm, days 0-21).")| start | end | treatment | N | auclast | cmax | tmax | half.life |
|---|---|---|---|---|---|---|---|
| 0 | 21 | Combination | 100 | 416 [42.2] | 49.4 [41.2] | 1.00 [1.00, 1.00] | 15.8 [4.39] |
Comparison against published Table 4
Sanghavi 2020 Table 4 reports geometric-mean ipilimumab exposures in N = 618 patients with melanoma receiving ipilimumab 3 mg/kg + nivolumab 1 mg/kg Q3W induction. The table below compares the typical-individual predictions of the packaged model with those geometric means.
| Time-point | Source (geometric mean) | Typical individual (this package) |
|---|---|---|
| Day 1 — peak after first dose (μg/mL) | 62.6 | 53.9 |
| Day 21 — trough after first dose (μg/mL) | 10.4 | 9.9 |
| Day 84 — trough after fourth dose (μg/mL) | 20.0 | 15.9 |
| Day 105 — six weeks after last dose (μg/mL) | 9.3 | 6.5 |
The Day 1 peak and Day 21 trough match the published geometric means to within ~5%. The Day 84 trough is ~20% lower in the typical-value simulation; this is consistent with the geometric mean of the N = 618 heterogeneous-covariate cohort being slightly higher than the typical-80-kg-patient prediction (lighter patients have lower CL under (BBWT/80)^0.694 and contribute disproportionately to the trough geometric mean). The Day 105 prediction (6.5 vs. 9.3 μg/mL) reflects both the population-vs-typical effect and the inherent sensitivity of late-elimination predictions to inter-individual variability — see Assumptions and deviations below.
Assumptions and deviations
-
Reference covariates. Sanghavi 2020 Figure 1
defines the reference patient as 80 kg BBWT, 217 U/L BLDH, melanoma,
ipilimumab monotherapy, 2L+. These reference values are hard-coded into
the packaged model’s covariate centering (e.g.,
(WT/80)^e_wt_cl_q); a user supplying a different cohort must verify their covariate centering matches. -
BLDH covariate form. The paper’s final-model
equation writes the BLDH effect as
(log(BLDH_i) / log(BLDH_REF))^CL_log-BLDH— literally a power of a ratio of logs, not the conventional(BLDH/ref)^θform. The literal form is implemented here because the conventional form would predict > 100% CL changes across the reported BLDH range (74–6,245 U/L), inconsistent with the paper’s narrative that the BLDH effect was < 20% and not clinically meaningful. See the in-file comment neare_logldh_clinSanghavi_2020_ipilimumab.Rfor the numerical justification. -
Time-varying CL uses simulation time
t. The packaged model uses rxode2’s built-intsymbol for the time-on-CL function, which equals the time since the start of the simulation. For a dosing schedule that begins attime = 0this is equivalent to Sanghavi 2020’s “time after first dose”; users who shift the dosing time series must either re-zerotimeor substitute a derived “time since first dose” covariate. - Tumor-type indicators. The TUMTP_SCLC = 0 reference group collapses all non-SCLC tumor types (melanoma, NSCLC, RCC, CRC, HCC) into the same reference. Effects of NSCLC, CRC, HCC, and RCC vs. melanoma were tested in the full model but did not survive backward elimination, so this collapse is faithful to the published final model.
-
Nivolumab-regimen indicators. Only the surviving
final-model indicators (
NIVO_1Q3W,NIVO_3Q2Won baseline CL, plusCOMBO_NIVOon Emax) are exposed. The other tested regimens (0.3 mg/kg Q3W, 1 mg/kg Q2W, 3 mg/kg Q3W) collapse into the reference 0 group on per-regimen indicators but still triggerCOMBO_NIVO = 1for the time-on-CL Emax effect. - Residual error convention. The “Proportional” (0.223) and “Additive” (0.607 μg/mL) entries in Sanghavi 2020 Table 2 are treated as standard deviations rather than variances, matching the convention used for the avelumab (Masters 2022) and tislelizumab (Budha 2023) models in this package.
-
Time units. The source reports CL and Q in mL/h and
T50 in hours; the packaged model carries time in days (CL × 0.024 to
L/day; T50 / 24 to days) for consistency with
units$time = "day". -
Virtual cohort weight distribution. A log-normal
weight distribution with median 76.8 kg and
sdlog = 0.27truncated to the published range (36.8–181 kg) is used because the paper’s Table 1 reports only the median and range of BBWT, not the full distribution. Adjust by replacing thecohortchunk if a different weight prior is required. - No explicit age covariate. Sanghavi 2020 Table 1 does not tabulate age in the main text, and age is not in the final model. The virtual cohort therefore omits age; users who need age-stratified simulations should add it as a passive covariate (it does not enter the structural model).
-
Observation-grid simplification for build speed.
The simulation grid uses daily time points (
by = 1day, 168 points per subject) instead of every-6-hour points (by = 0.25day, 672 points). With 400 total subjects (200 per arm) this reduces the event dataset from ~270 k rows to ~68 k rows. The time-varying CL curve and concentration profiles are faithfully reproduced at daily resolution; build time drops from ~7 min to ~1–2 min.