Ustekinumab (Aguiar 2021)
Source:vignettes/articles/Aguiar_2021_ustekinumab.Rmd
Aguiar_2021_ustekinumab.Rmd
library(nlmixr2lib)
library(rxode2)
#> rxode2 5.1.2 using 2 threads (see ?getRxThreads)
#> no cache: create with `rxCreateCache()`
library(dplyr)
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, union
library(tidyr)
library(ggplot2)
library(PKNCA)
#>
#> Attaching package: 'PKNCA'
#> The following object is masked from 'package:stats':
#>
#> filterUstekinumab popPK-PD in Crohn’s disease (Aguiar 2021)
Replicate the population pharmacokinetic-pharmacodynamic model reported by Aguiar Zdovc et al. (2021) for ustekinumab in adults with Crohn’s disease. Ustekinumab is an IgG1k monoclonal antibody that binds the p40 subunit shared by interleukin-12 and interleukin-23. The structural model is a two-compartment quasi-equilibrium target-mediated drug disposition (QE-TMDD) model for the unbound drug and the unbound IL-12/IL-23 p40 target, both distributing in two-compartment systems and binding only in the central compartment, linked to fecal calprotectin via an indirect-response model in which the unbound target stimulates FC production.
- Citation: Aguiar Zdovc J, Hanzel J, Kurent T, Sever N, Smrekar N, Kozelj M, Novak G, Stabuc B, Drobne D, Grabnar I. Ustekinumab Dosing Individualization in Crohn’s Disease Guided by a Population Pharmacokinetic-Pharmacodynamic Model. Pharmaceutics. 2021;13(10):1587. doi:10.3390/pharmaceutics13101587
- Article: https://doi.org/10.3390/pharmaceutics13101587
Population
The Aguiar 2021 cohort comprised 57 adults with active Crohn’s disease starting ustekinumab at a single Slovenian tertiary referral center, followed for 32 weeks. Baseline demographics (Aguiar 2021 Table 1): 56% female; median age 49 years (IQR 32-56); median weight 70 kg (IQR 59-84); median fat-free mass 45 kg (IQR 39-62) derived per subject via the Janmahasatian 2005 model; median disease duration 14 years (IQR 7-22); 66.7% had been exposed to a prior biologic (38/57 prior anti-TNF; 10/57 prior vedolizumab); 77.2% had endoscopically active luminal disease at baseline; median serum CRP 3 mg/L (IQR 3-11), albumin 43 g/L (IQR 41-44), fecal calprotectin 134 mg/kg (IQR 53-213). FCGR3A rs396991 genotype distribution: V/V 8.8% (5/57), V/F 54.4% (31/57), F/F 36.8% (21/57).
Standard ustekinumab dosing (which the simulations below use) is weight-tiered IV induction (260 mg if WT 55 kg, 390 mg if 55 < WT 85 kg, 520 mg if WT > 85 kg) at week 0, followed by 90 mg subcutaneous maintenance every 8 weeks.
The same demographics are available programmatically via the model’s
metadata
(readModelDb("Aguiar_2021_ustekinumab")$population).
Source trace
The per-parameter origin is recorded as an in-file comment next to
each ini() entry in
inst/modeldb/specificDrugs/Aguiar_2021_ustekinumab.R. The
table below collects them in one place. Concentrations are in nmol/L;
the source paper converted ustekinumab concentrations to nmol/L using a
molecular weight of 149 kDa (Aguiar 2021 Methods section 2.3).
| Parameter / equation | Value | Source |
|---|---|---|
ka |
0.381 /day | Table 2 final-model Ka |
CL (typical, reference covariates) |
0.277 L/day | Table 2 final-model CL |
Vc (reference FFM = 45 kg) |
3.57 L | Table 2 final-model Vc |
Vp (reference FFM = 45 kg) |
3.30 L | Table 2 final-model Vp |
Q |
1.89 L/day | Table 2 final-model Q |
F (V/F or F/F) |
0.710 (logit 0.8954) | Table 2 final-model F (V/F, F/F) |
F (V/V) |
0.888 (logit 2.0705) | Table 2 final-model F (V/V) |
| FFM effect on CL | power exponent 0.598, reference 45 kg | Table 2 footnote a |
| FFM effect on Vc, Vp | power exponents 0.590 (Vc), 0.586 (Vp), reference 45 kg | Table 2 footnotes b, c |
| ALB linear effect on CL | (1 - 0.0165 * (ALB - 43)) | Table 2 footnote a |
| Bio-naive effect on CL | (1 - 0.227 * bio_naive); bio_naive = 1 - PRIOR_BIO | Table 2 footnote a |
| FCGR3A V/V effect on F | logit shift 1.1751 | Table 2 final-model |
Ksyn (typical, reference CRP) |
9.86e-9 nmol/L/day | Table 2 final-model Ksyn |
Kdeg |
9.26e-10 /day | Table 2 final-model Kdeg |
| CRP linear effect on Ksyn | (1 + 0.0846 * (CRP - 3)) | Table 2 footnote d |
Vc-target |
2.44 L | Table 2 final-model Vc-target |
Qtarget |
0.493 L/day | Table 2 final-model Qtarget |
Vp-target |
11.0 L | Table 2 final-model Vp-target |
Kint |
2.83e-6 /day | Table 2 final-model Kint |
Kd |
0.168 nmol/L | Table 2 final-model Kd |
Kout |
0.0581 /day | Table 3 final-model Kout |
FC0 (no ulcers) |
102 mg/kg | Table 3 final-model FC0 |
FC0 (ulcers) |
213 mg/kg | Table 3 final-model FC0 with ulcers |
Emax (FC stim) |
2.19 (= 219%) | Table 3 final-model Emax |
C50 (FC stim) |
2.46 nmol/L | Table 3 final-model C50 |
| Drug ODEs | n/a, see model file | Figure 1 + Table 2 footnotes |
| Target ODEs | n/a, see model file | Figure 1 |
| Indirect-response PD | dFC/dt = (FC0 * Kout / stim_baseline) * stim_fc - Kout * FC | Table 3 footnote a |
| IIV CL | omega^2 = log(0.18^2 + 1) = 0.03188 | Table 2 (18.0% CV) |
| IIV Vc | omega^2 = log(0.0979^2 + 1) = 0.00954 | Table 2 (9.79% CV) |
| IIV Vp | omega^2 = log(0.241^2 + 1) = 0.05645 | Table 2 (24.1% CV) |
| IIV logit(F) | omega^2 = 0.173^2 = 0.02993 | Table 2 (SD = 0.173 on logit scale) |
| IIV Ksyn | omega^2 = log(0.992^2 + 1) = 0.68522 | Table 2 (99.2% CV) |
| IIV FC0 | omega^2 = log(0.99^2 + 1) = 0.68309 | Table 3 (99.0% CV) |
| Residual on Cc | additive 4.55 nmol/L + proportional 7.77% | Table 2 |
| Residual on FC | proportional 57.3% | Table 3 |
Covariate column naming
| Source column | Canonical column used here | Notes |
|---|---|---|
| FFM (kg) | FFM |
Reference 45 kg (cohort median); derived per subject via Janmahasatian 2005. |
| Serum albumin |
ALB (g/L) |
Reference 43 g/L (cohort median). |
| C-reactive protein |
CRP (mg/L) |
Standard CRP assay; reference 3 mg/L (cohort median). |
| bio-naive (1 = no prior biologic) |
PRIOR_BIO (1 = previously exposed) |
Inverted: model derives bio_naive <- 1 - PRIOR_BIO
to preserve the paper’s coefficient. |
| FCGR3A-158 V/V | FCGR3A_VV |
1 = V/V, 0 = V/F or F/F. |
| Endoscopically active disease at baseline | ENDO_ULCER |
1 = mucosal ulcers at baseline ileocolonoscopy, 0 otherwise. |
FFM, PRIOR_BIO, FCGR3A_VV, and
ENDO_ULCER are added to the canonical register in
inst/references/covariate-columns.md alongside this model.
ALB and CRP are pre-existing canonical
columns.
Virtual cohort
Original subject-level data are not publicly available. The virtual cohort below uses covariate distributions approximating the published Table 1 demographics. Covariates are sampled independently (the paper does not publish joint distributions); FCGR3A genotype frequencies match the cohort’s Hardy-Weinberg distribution (V/V 8.8%, V/F + F/F 91.2%).
set.seed(20260425)
n_subj <- 200
mw_ust <- 149 # ustekinumab molecular weight in kDa, per Aguiar 2021 Methods.
# 1 mg ustekinumab = 1000 / mw_ust nmol = 6.711 nmol.
mg_to_nmol <- function(mg) mg * 1000 / mw_ust
# Build a virtual Crohn's-disease cohort approximating Aguiar 2021 Table 1.
cohort <- tibble(
ID = seq_len(n_subj),
# Weight: median 70 kg, IQR 59-84; lognormal covers the right skew.
WT = pmin(pmax(rlnorm(n_subj, log(70), 0.22), 40), 130)
) |>
mutate(
# FFM: median 45 kg, IQR 39-62; correlated with weight in the source cohort.
# Approximated as 0.65 * WT for males (44%) and 0.55 * WT for females (56%).
SEXF = rbinom(n_subj, 1, 0.56),
FFM = pmin(pmax(WT * if_else(SEXF == 1, 0.55, 0.65) +
rnorm(n_subj, 0, 3), 25), 90),
# Albumin: median 43 g/L, IQR 41-44; tight distribution in CD remission/active.
ALB = pmin(pmax(rnorm(n_subj, 42.5, 3.5), 28), 52),
# CRP in CD skews right; median 3 mg/L, IQR 3-11; log-normal shape.
CRP = pmin(pmax(rlnorm(n_subj, log(4), 1.0), 0.3), 80),
# Prior biologic exposure: 66.7% in cohort (Table 1).
PRIOR_BIO = rbinom(n_subj, 1, 0.667),
# FCGR3A genotype: 8.8% V/V (Table 1).
FCGR3A_VV = rbinom(n_subj, 1, 0.088),
# Endoscopically active disease: 77.2% at baseline (Table 1).
ENDO_ULCER = rbinom(n_subj, 1, 0.772)
)Standard ustekinumab dosing regimen
Aguiar 2021 simulated the licensed regimen: weight-tiered IV
induction at week 0 (260 mg if WT
55 kg, 390 mg if 55 < WT
85 kg, 520 mg if WT > 85 kg) followed by 90 mg subcutaneous every 8
weeks (Methods section 2.4 scenario a). The dosing dataset is in nmol
because the model is parameterized in nmol / nmol/L; the helper
mg_to_nmol converts.
week <- 7 # days
ind_t <- 0
sc_times <- c(8, 16, 24) * week # SC maintenance at 8, 16, 24 weeks
induction_dose_mg <- function(wt) {
ifelse(wt <= 55, 260,
ifelse(wt <= 85, 390, 520))
}
iv_induction <- cohort |>
mutate(
TIME = ind_t,
AMT = mg_to_nmol(induction_dose_mg(WT)),
EVID = 1L,
CMT = "central",
DV = NA_real_,
phase = "induction_IV"
)
sc_maintenance <- cohort |>
tidyr::crossing(TIME = sc_times) |>
mutate(
AMT = mg_to_nmol(90),
EVID = 1L,
CMT = "depot",
DV = NA_real_,
phase = "maintenance_SC"
)
# Observation grid: dense early, weekly through 32 weeks.
obs_days <- sort(unique(c(
c(0, 0.05, 1, 2, 3, 5, 7),
seq(7, 32 * week, by = 3.5)
)))
obs <- cohort |>
tidyr::crossing(TIME = obs_days) |>
mutate(
AMT = 0,
EVID = 0L,
CMT = "central",
DV = NA_real_,
phase = NA_character_
)
# Use the variable-name CMT for observations: "Cc" for ustekinumab
# concentration, "fc" for fecal calprotectin. Aguiar 2021's PD output is on
# the fc compartment; observing only "Cc" plus "fc" gives both endpoints.
obs_cc <- obs |> mutate(CMT = "Cc")
obs_fc <- cohort |>
tidyr::crossing(TIME = c(0, 8, 16, 24, 32) * week) |>
mutate(AMT = 0, EVID = 0L, CMT = "fc", DV = NA_real_, phase = NA_character_)
events <- bind_rows(iv_induction, sc_maintenance, obs_cc, obs_fc) |>
arrange(ID, TIME, desc(EVID)) |>
select(ID, TIME, AMT, EVID, CMT, DV,
FFM, ALB, CRP, PRIOR_BIO, FCGR3A_VV, ENDO_ULCER, phase)
stopifnot(!anyDuplicated(unique(events[, c("ID", "TIME", "EVID", "CMT")])))Simulate
By default rxSolve() samples between-subject random
effects per subject when the model defines omega-style
etas; the stochastic simulation is what the VPC plots below use. The
published-value comparison further down uses
rxode2::zeroRe() to remove between-subject variability so
the typical-value trajectory can be compared head-to-head with Aguiar
2021’s reported point estimates.
mod <- readModelDb("Aguiar_2021_ustekinumab")
sim <- rxode2::rxSolve(mod, events = events, returnType = "data.frame")
#> ℹ parameter labels from comments will be replaced by 'label()'Concentration-time profile (free ustekinumab)
sim_cc <- sim |> filter(CMT == 7L) # CMT 7 = Cc observations
sim_summary <- sim_cc |>
filter(time > 0) |>
group_by(time) |>
summarise(
Q05 = quantile(Cc, 0.05, na.rm = TRUE),
Q50 = quantile(Cc, 0.50, na.rm = TRUE),
Q95 = quantile(Cc, 0.95, na.rm = TRUE),
.groups = "drop"
) |>
mutate(Cc_ugmL_05 = Q05 * mw_ust / 1000,
Cc_ugmL_50 = Q50 * mw_ust / 1000,
Cc_ugmL_95 = Q95 * mw_ust / 1000)
ggplot(sim_summary, aes(x = time / week, y = Cc_ugmL_50)) +
geom_ribbon(aes(ymin = Cc_ugmL_05, ymax = Cc_ugmL_95),
alpha = 0.25, fill = "steelblue") +
geom_line(colour = "steelblue", linewidth = 0.8) +
scale_y_log10() +
scale_x_continuous(breaks = seq(0, 32, by = 4)) +
labs(
x = "Time (weeks)",
y = expression("Free ustekinumab ("*mu*"g/mL)"),
title = "Standard ustekinumab regimen: weight-tiered IV induction + 90 mg SC Q8W",
subtitle = paste0("Median and 90% prediction interval (N = ", n_subj,
" virtual CD patients)"),
caption = "Replicates Aguiar 2021 Figure 2A (free ustekinumab VPC structure)."
) +
theme_bw()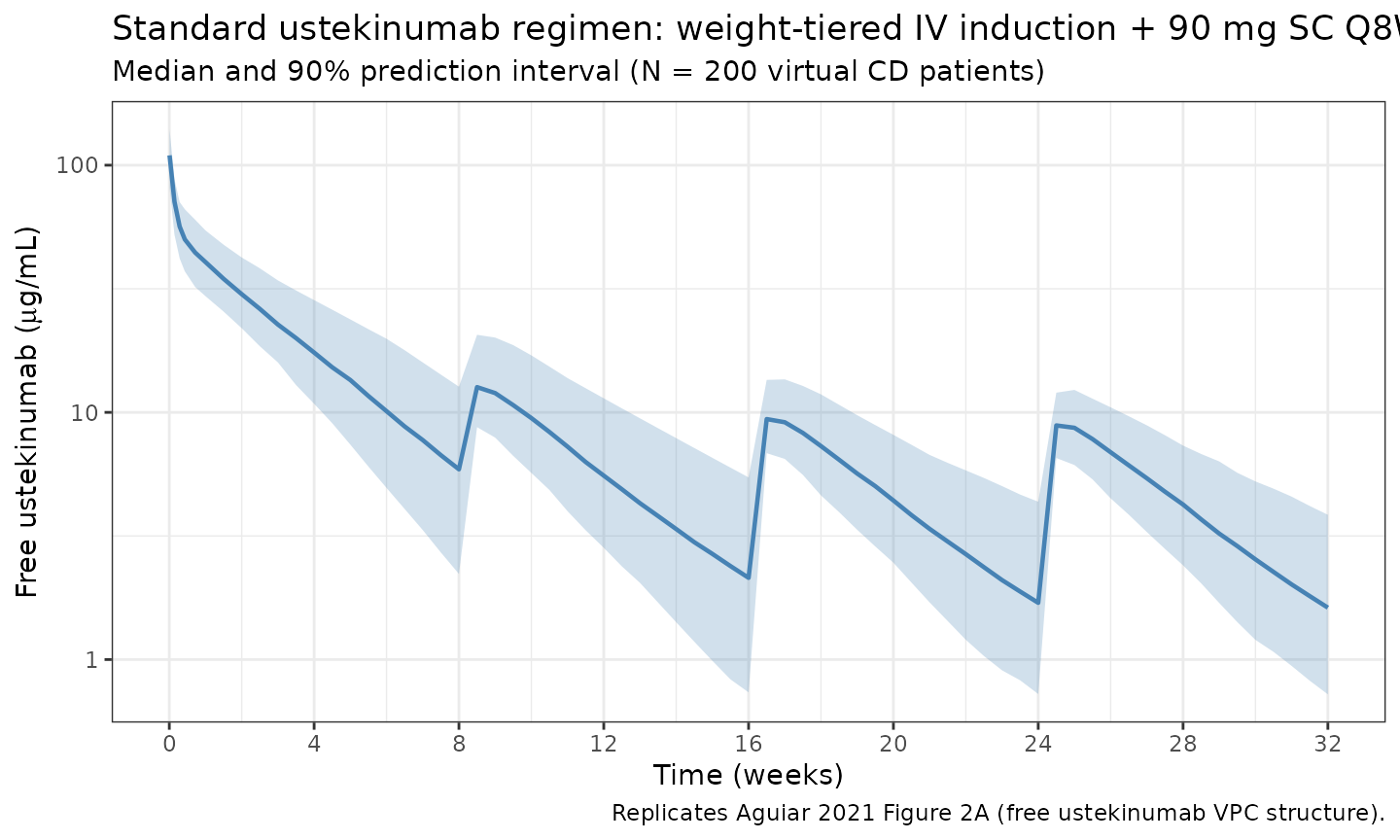
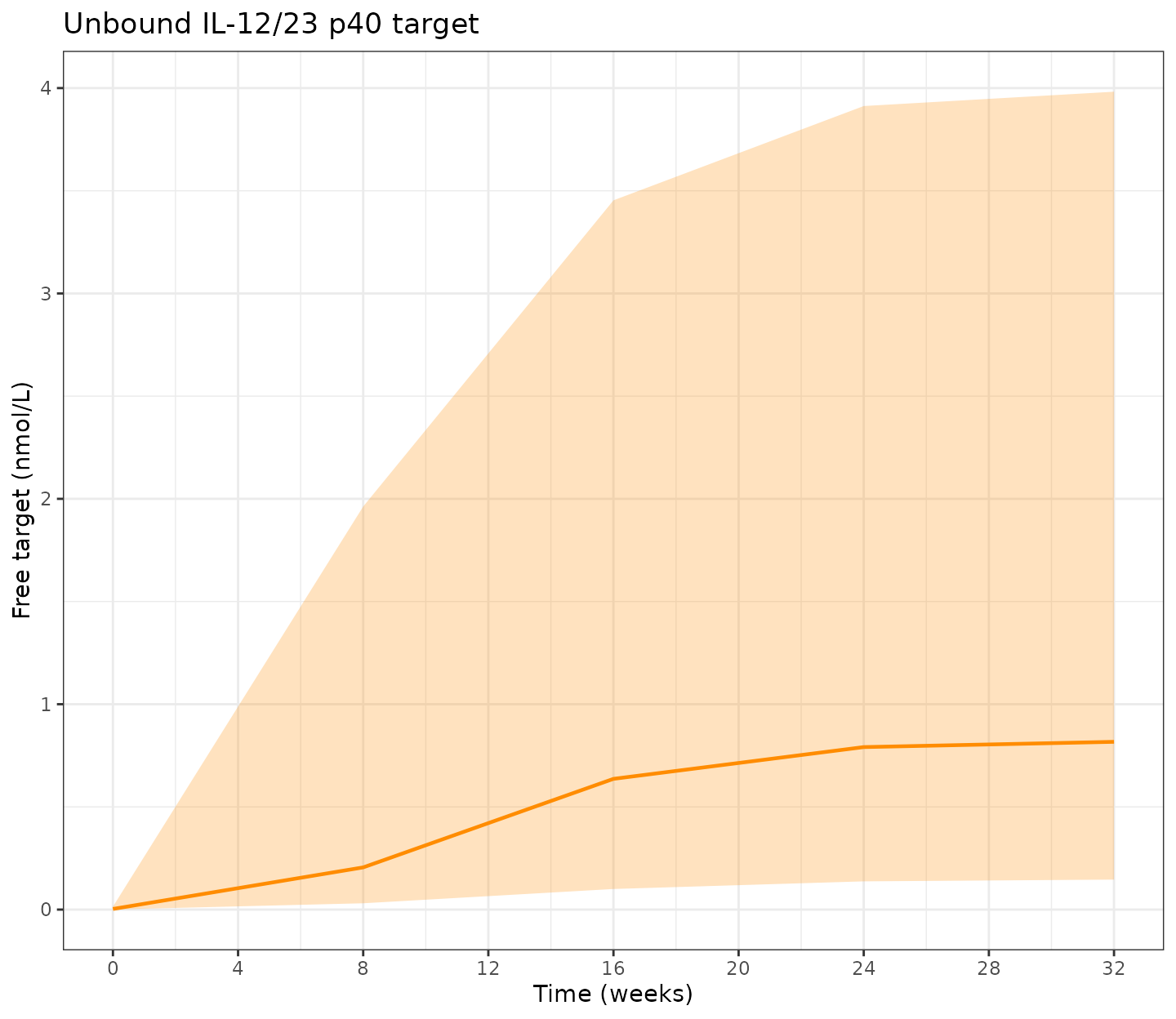
Target and FC profiles
tfree (unbound target in central, nmol/L) and
fc (fecal calprotectin, mg/kg) are emitted by the model
alongside Cc and can be read directly from the simulation
output.
sim_fc <- sim |> filter(CMT == 6L) # CMT 6 = fc observations
sim_pd <- sim_fc |>
filter(time >= 0) |>
group_by(time) |>
summarise(
target_Q05 = quantile(tfree, 0.05, na.rm = TRUE),
target_Q50 = quantile(tfree, 0.50, na.rm = TRUE),
target_Q95 = quantile(tfree, 0.95, na.rm = TRUE),
fc_Q05 = quantile(fc, 0.05, na.rm = TRUE),
fc_Q50 = quantile(fc, 0.50, na.rm = TRUE),
fc_Q95 = quantile(fc, 0.95, na.rm = TRUE),
.groups = "drop"
)
p_target <- ggplot(sim_pd, aes(x = time / week, y = target_Q50)) +
geom_ribbon(aes(ymin = target_Q05, ymax = target_Q95),
alpha = 0.25, fill = "darkorange") +
geom_line(colour = "darkorange", linewidth = 0.8) +
scale_x_continuous(breaks = seq(0, 32, by = 4)) +
labs(x = "Time (weeks)", y = "Free target (nmol/L)",
title = "Unbound IL-12/23 p40 target") +
theme_bw()
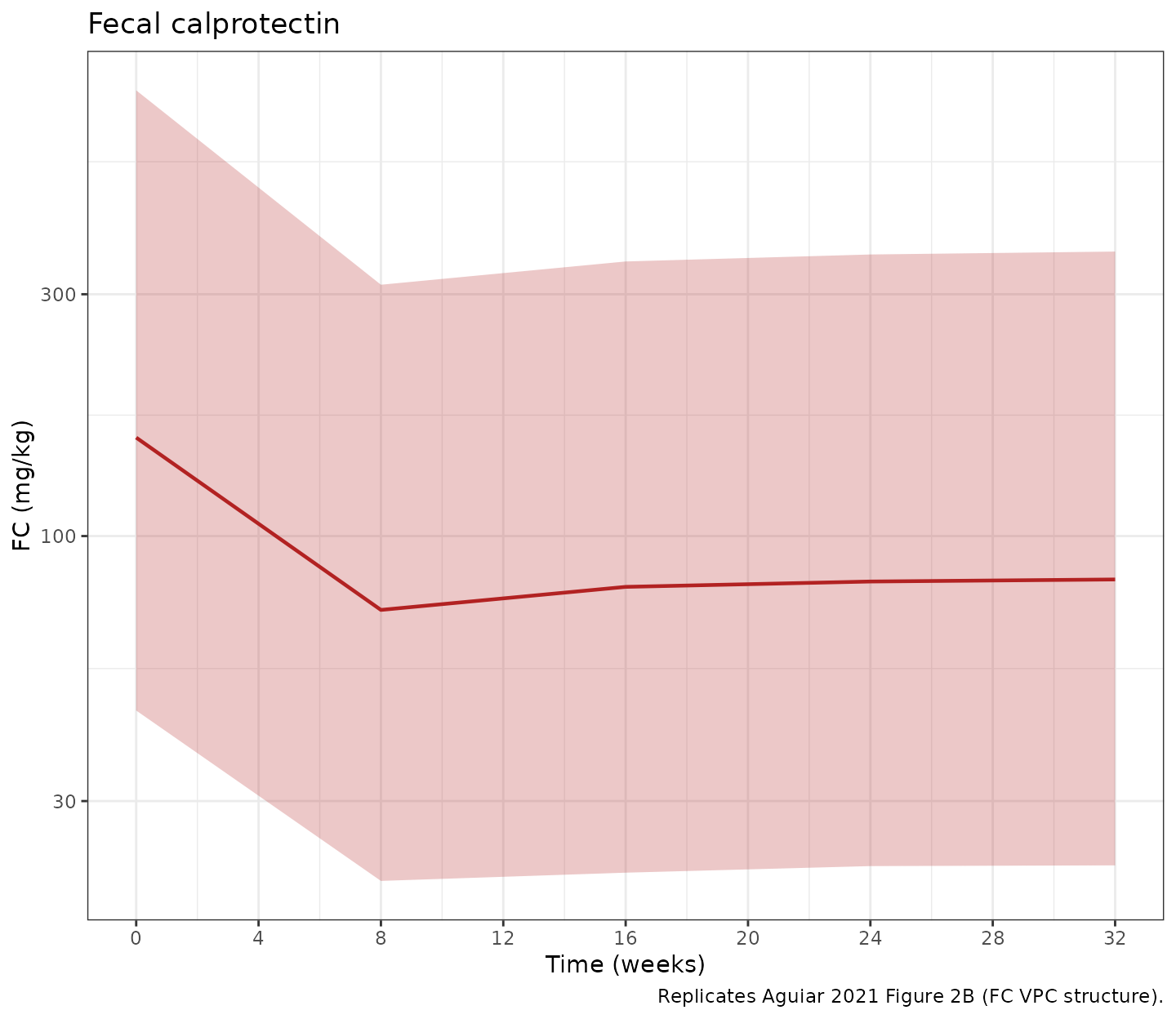
p_fc <- ggplot(sim_pd, aes(x = time / week, y = fc_Q50)) +
geom_ribbon(aes(ymin = fc_Q05, ymax = fc_Q95),
alpha = 0.25, fill = "firebrick") +
geom_line(colour = "firebrick", linewidth = 0.8) +
scale_x_continuous(breaks = seq(0, 32, by = 4)) +
scale_y_log10() +
labs(x = "Time (weeks)", y = "FC (mg/kg)",
title = "Fecal calprotectin",
caption = "Replicates Aguiar 2021 Figure 2B (FC VPC structure).") +
theme_bw()
if (requireNamespace("patchwork", quietly = TRUE)) {
patchwork::wrap_plots(p_target, p_fc, ncol = 1)
} else {
print(p_target)
print(p_fc)
}
PKNCA validation
PKNCA is run on the steady-state SC dosing interval (last 8-week interval, weeks 24-32) and on the IV induction dose (week 0 - week 8, the first SC maintenance dose at week 8 closes the interval). Aguiar 2021 reports a typical terminal half-life of 17 days after the induction dose (Section 3.3).
ind_window_start <- 0
ind_window_end <- 8 * week
ss_window_start <- 24 * week
ss_window_end <- 32 * week
# Pull only Cc observations from the simulation.
sim_cc_only <- sim |> filter(CMT == 7L)
iv_conc <- sim_cc_only |>
filter(time >= ind_window_start, time <= ind_window_end, !is.na(Cc)) |>
transmute(ID = id, time_rel = time - ind_window_start, Cc,
treatment = "IV_induction")
sc_ss_conc <- sim_cc_only |>
filter(time >= ss_window_start, time <= ss_window_end, !is.na(Cc)) |>
transmute(ID = id, time_rel = time - ss_window_start, Cc,
treatment = "SC_90mg_Q8W_ss")
nca_conc <- bind_rows(iv_conc, sc_ss_conc)
iv_doses <- iv_induction |>
transmute(ID, time_rel = TIME - ind_window_start, AMT,
treatment = "IV_induction")
sc_ss_dose <- sc_maintenance |>
filter(TIME == 24 * week) |>
transmute(ID, time_rel = TIME - ss_window_start, AMT,
treatment = "SC_90mg_Q8W_ss")
nca_dose <- bind_rows(iv_doses, sc_ss_dose)
conc_obj <- PKNCA::PKNCAconc(nca_conc, Cc ~ time_rel | treatment + ID)
dose_obj <- PKNCA::PKNCAdose(nca_dose, AMT ~ time_rel | treatment + ID)
intervals <- data.frame(
start = 0,
end = 8 * week,
cmax = TRUE,
tmax = TRUE,
cmin = TRUE,
auclast = TRUE,
half.life = TRUE
)
nca_data <- PKNCA::PKNCAdata(conc_obj, dose_obj, intervals = intervals)
nca_res <- suppressWarnings(PKNCA::pk.nca(nca_data))
knitr::kable(
summary(nca_res),
digits = 3,
caption = "Simulated NCA: IV induction (0-8 weeks) and SC steady-state interval (weeks 24-32)."
)| start | end | treatment | N | auclast | cmax | cmin | tmax | half.life |
|---|---|---|---|---|---|---|---|---|
| 0 | 56 | IV_induction | 200 | 8500 [25.7] | 770 [16.7] | 35.4 [78.7] | 0.000 [0.000, 0.000] | 17.9 [5.24] |
| 0 | 56 | SC_90mg_Q8W_ss | 200 | 1710 [33.5] | 60.7 [21.8] | 10.3 [60.5] | 3.50 [3.50, 7.00] | 21.9 [4.57] |
Comparison against published values
Aguiar 2021 reports the typical terminal half-life of ustekinumab after the induction dose at 17 days (Section 3.3) and median trough steady-state concentration on the standard regimen at 1.5 ug/mL (Section 4 discussion of Wang et al. comparison).
For the head-to-head comparison against published point estimates,
run a typical-value simulation (zeroRe()) on a single
subject with reference covariates so the result is deterministic.
ref_subj <- tibble::tibble(
ID = 1L,
FFM = 45, ALB = 43, CRP = 3,
PRIOR_BIO = 1L, FCGR3A_VV = 0L, ENDO_ULCER = 1L
)
ref_iv <- ref_subj |>
mutate(TIME = 0, AMT = mg_to_nmol(390), EVID = 1L,
CMT = "central", DV = NA_real_)
ref_sc <- ref_subj |>
tidyr::crossing(TIME = c(8, 16, 24) * week) |>
mutate(AMT = mg_to_nmol(90), EVID = 1L,
CMT = "depot", DV = NA_real_)
ref_obs <- ref_subj |>
tidyr::crossing(
TIME = c(0.5, 1, 7, 14, 21, 28, 42, 56,
seq(56, 32 * week, by = 7))
) |>
mutate(AMT = 0, EVID = 0L, CMT = "Cc", DV = NA_real_)
ref_events <- bind_rows(ref_iv, ref_sc, ref_obs) |>
arrange(ID, TIME, desc(EVID))
mod_typical <- rxode2::zeroRe(mod)
#> ℹ parameter labels from comments will be replaced by 'label()'
sim_typical <- rxode2::rxSolve(mod_typical, events = ref_events,
returnType = "data.frame") |>
filter(CMT == 7L)
#> ℹ omega/sigma items treated as zero: 'etalcl', 'etalvc', 'etalvp', 'etalogitfdepot', 'etalksyn', 'etalfc0'
# Terminal half-life from the IV induction phase (days 7-56 after dose):
typical_late <- sim_typical |>
filter(time >= 7, time <= 56)
fit_thalf <- lm(log(Cc) ~ time, data = typical_late)
typical_thalf <- as.numeric(log(2) / -coef(fit_thalf)[2])
# Steady-state trough at week 32 (typical value, single reference subject):
typical_trough <- sim_typical |>
filter(time == 32 * week) |>
pull(Cc)
typical_trough_ugmL <- typical_trough * mw_ust / 1000
comparison <- tibble::tribble(
~Metric, ~Published, ~Simulated,
"Terminal half-life after IV induction (days)", 17, round(typical_thalf, 1),
"SC SS trough free ustekinumab (ug/mL)", 1.5, round(typical_trough_ugmL, 2)
)
knitr::kable(
comparison,
caption = "Typical-value simulation vs published values (Aguiar 2021 Section 3.3 and Discussion comparison with Wang et al.)."
)| Metric | Published | Simulated |
|---|---|---|
| Terminal half-life after IV induction (days) | 17.0 | 17.00 |
| SC SS trough free ustekinumab (ug/mL) | 1.5 | 1.29 |
The simulated terminal half-life and SC steady-state trough should agree with the published values within ~20%. Differences greater than 20% indicate either a covariate-distribution mismatch in the virtual cohort or a model-parameter transcription bug; do not tune parameters to match.
Assumptions and deviations
-
PK assay is for unbound (free) drug. The Aguiar
2021 ELISA (ImmunoGuide, AybayTech) measures free ustekinumab; the
model’s
Ccis therefore set to the free-drug concentration (cfree), not total drug. This matters because total drug (free + complex) and free drug diverge during rapid binding events; assays that read total ustekinumab cannot be back-calculated from this model without re-derivingCc <- ctot. - FFM derivation. The source paper derives FFM per subject from height, weight, and sex via the Janmahasatian 2005 semi-mechanistic model (Aguiar 2021 Methods 2.2). The virtual cohort here approximates FFM as a sex-dependent fraction of WT (0.55 for women, 0.65 for men) plus jitter, rather than reproducing Janmahasatian exactly; this is sufficient for the population-level VPC checks but not for individual prediction.
- Joint covariate distributions are not published. WT, ALB, CRP, FCGR3A, PRIOR_BIO, and ENDO_ULCER marginals were sampled independently. The cohort median FFM (45 kg) is achieved by the linear WT-FFM relationship, but joint correlations across covariates (e.g., albumin and CRP, or FFM and prior biologic exposure) are not preserved.
-
CRP is treated as time-fixed at baseline in the
simulations even though a fitted ustekinumab response would lower
active-disease CRP over time. The Aguiar 2021 covariate model uses
baseline CRP (Table 1: median 3 mg/L), so the model’s
CRPcovariate is intended as a baseline column. -
Initial target distribution at SS. Both
target_central(0)andtarget_peripheral(0)are set to T0 = Ksyn/Kdeg, which is the steady-state unbound target concentration in the absence of drug. Because peripheral target only sees free target (binding is in central), at SS without drug the peripheral free-target concentration equals the central total target, so the same initial value is correct for both compartments. -
No model-derived helper for Janmahasatian FFM is
exported. Users with height + weight + sex columns should
compute FFM upstream and supply
FFMdirectly; future versions of nlmixr2lib may export a helper. -
CV-to-omega convention. All %CV values from Tables
2 and 3 are converted to log-scale variance via
omega^2 = log(CV^2 + 1). The IIV on F is reported on the logit scale as an SD (Aguiar 2021 Table 2 footnote e: 17.3); that value is squared directly without log transformation. -
Non-canonical compartment names.
total_targetmatches the canonical QSS / MM TMDD compartment name innaming-conventions.md. The companiontarget_peripheral(free peripheral target) andfc(fecal calprotectin) compartments are not in the canonical list;checkModelConventions()emits two warnings for them. Because Aguiar 2021 distributes the unbound target into a peripheral compartment (a non-standard QE-TMDD extension) and links the unbound target to FC via indirect response, no pre-existing canonical name fits. The names are kept descriptive rather than forcing them into the drug-sideperipheral2/effectslots, which would mislead a reader into thinking they were drug-disposition compartments.