Model and source
- Citation: Hood T, Slob CJ, Slager J, Yates JWT, Bouzom F, Mistry HB. Pharmacokinetic-Pharmacodynamic Modelling of Systemic IL13 Blockade by Monoclonal Antibody Therapy: A Free Assay Disguised as Total. Pharmaceutics. 2021;13(5):613.
- Article (open access): doi:10.3390/pharmaceutics13050613
- Trial registry: NCT02388347
MEDI7836 is a fully human IgG1λ-YTE anti-IL13 monoclonal antibody developed as a back-up to tralokinumab/lebrikizumab. The study was a phase-1 randomized placebo-controlled single-ascending-dose trial in healthy adult males. The novel scientific contribution of the paper is not the PK structure itself (a standard 2-compartment SC model) but the integrated PK-PD interpretation: although the IL13 immunoassay was validated as a “free IL13” assay, the observed serum IL13 increased after dosing rather than decreasing, and the authors show that the data are well described if the assay also captures ~4% of the bound MEDI7836:IL13 complex. The clinical programme was halted because the model-projected efficacious dose range did not match the best-in-class target product profile.
Population
Hood 2021 Table 1 summarises the trial cohort:
- N: 32 healthy adult males (24 active, 8 placebo).
- Sex: 100% male (study inclusion criterion).
- Age: median 35 years overall (range 18-50).
- Body weight: median 76.5 kg overall (IQR 60.4-95.2).
- Race: White 19/32 (59.4%), Black/African American 8/32 (25.0%), Asian 5/32 (15.6%).
- Dose levels: 30, 105, 300, or 600 mg single SC dose, with six active subjects per cohort and eight placebo subjects shared across cohorts.
- Region: United Kingdom, single site.
- Sampling: average 13.9 PK samples per active subject and 14.9 PD (IL13) samples per subject across the 281-day follow-up window.
- Immunogenicity: 21/24 active subjects became ADA-positive post-baseline (87.5%); 16/24 (67%) classified persistent ADA-positive.
Two subjects whose PD profiles appeared swapped in the data (one nominally placebo subject with an active-like IL13 response, one nominally 600 mg subject with a placebo-like response) were excluded from the PD analysis dataset.
The same information is available programmatically via
readModelDb("Hood_2021_medi7836")$population.
Source trace
Per-parameter provenance is recorded next to each ini()
entry in inst/modeldb/specificDrugs/Hood_2021_medi7836.R.
The table below collects them in one place.
| Element | Source | Value / form |
|---|---|---|
| Ka | Hood 2021 Table 2 | 0.156 1/day [0.137-0.183] |
| CL/F | Hood 2021 Table 2 | 0.441 L/day [0.366-0.587] (ADA-negative reference) |
| V2/F | Hood 2021 Table 2 | 2.83 L [2.21-4.06] (free MEDI7836 central) |
| V3/F | Hood 2021 Table 2 | 8.03 L [6.82-9.52] (peripheral; shared with complex) |
| Q/F | Hood 2021 Table 2 | 0.825 L/day [0.638-1.13] (shared with complex) |
| Vcx/F | Hood 2021 Table 2 | 13.6 L [10.5-16.7] (IL13:MEDI7836 complex central) |
| Kin | Hood 2021 Table 2 | 0.0173 [0.0136-0.0227] (paper labels nmol/d; the math implies nM/day at the implicit 1-L IL13 reference volume — see Assumptions) |
| Kout | Hood 2021 Table 2 | 180 1/day [143-227] |
| Cx fraction | Hood 2021 Table 2 | 4.29% [2.98-5.96%] (assay fraction of complex) |
| Kon (fixed) | Hood 2021 Section 2.5 | 138.24 1/(nM·day), Biacore in-vitro |
| Koff (fixed) | Hood 2021 Section 2.5 | 0.69 1/day, Biacore in-vitro |
| ADA on CL | Hood 2021 Table 2 (Eq. 5) |
(1 + 0.717 * ADA_POS), ADA-positive post-baseline |
| IIV CL/F | Hood 2021 Table 2 | CV 53.3% → ω² = log(1 + 0.533²) = 0.250303 |
| IIV V2/F | Hood 2021 Table 2 | CV 72.7% → ω² = 0.424439 |
| IIV V3/F | Hood 2021 Table 2 | CV 46.2% → ω² = 0.193389 |
| IIV Q/F | Hood 2021 Table 2 | CV 69.6% → ω² = 0.395029 |
| IIV Kout | Hood 2021 Table 2 | CV 57.1% → ω² = 0.282271 |
| IIV Cx fraction | Hood 2021 Table 2 | CV 154% → ω² = 1.215281 |
| PK residual additive (ε1) | Hood 2021 Table 2 | 0.0546 ng/mL = 0.0546 µg/L |
| PK residual proportional (ε2) | Hood 2021 Table 2 | 13.0% |
| PD residual (ε3, on log scale) | Hood 2021 Table 2 | 6.9% |
| PK observation form | Hood 2021 Eq. 2 | combined: Cc = pred * (1 + ε2) + ε1
|
| PD observation form | Hood 2021 Eq. 3 | log-scale, proportional:
pdIL13 = log(IL13 + FR * complex_central) * (1 + ε3)
|
Covariate column naming
| Source column | Canonical column used here | Notes |
|---|---|---|
| ADA | ADA_POS |
Time-fixed binary, 1 = ADA-positive at any post-baseline visit. The paper’s “ADA-positive post-baseline” classification was carried as a time-fixed covariate; the authors discuss the time-invariance assumption as a limitation that may have biased the magnitude of the effect. |
Body weight, age, race, and pre-existing ADA were all tested but not retained in the final model (Hood 2021 Section 3.1).
Virtual cohort
Original observed concentrations are not publicly available. The figures below use a virtual population whose dose levels match the trial design (30, 105, 300, 600 mg SC). For the typical-value replications (Figures 2, 3, 7) we zero out IIV; for the VPC-style replication of Figure 5 we sample IIV from the published log-normal distributions. ADA status is held at 0 in the typical-value cohort and varied in the VPC cohort.
set.seed(20210601)
doses <- c(30, 105, 300, 600)
n_per_dose <- 60L # virtual cohort -- larger than the trial for smoother percentiles
# Two dosing-event-only frames: one per-subject for typical-value sims
# (24 subjects, 6 per active dose level), and a larger one for stochastic VPC.
make_cohort <- function(n, dose, ada_pct = 87.5, id_offset = 0L) {
n_ada_pos <- round(n * ada_pct / 100)
ada_vec <- c(rep(1L, n_ada_pos), rep(0L, n - n_ada_pos))
tibble(
id = id_offset + seq_len(n),
dose = dose,
ADA_POS = sample(ada_vec)
)
}
# Typical-value cohort: 6 subjects per dose, ADA-negative (paper Figure 2 was
# typical-prediction overlays, so use the ADA-negative reference parameter set).
typical_subjects <- bind_rows(
make_cohort(6, 30, ada_pct = 0, id_offset = 0L),
make_cohort(6, 105, ada_pct = 0, id_offset = 100L),
make_cohort(6, 300, ada_pct = 0, id_offset = 200L),
make_cohort(6, 600, ada_pct = 0, id_offset = 300L)
)
# Stochastic cohort: n_per_dose subjects per dose, 87.5% ADA-positive (matches
# Hood 2021 Table 1 active-cohort post-baseline rate).
stoch_subjects <- bind_rows(
make_cohort(n_per_dose, 30, id_offset = 1000L),
make_cohort(n_per_dose, 105, id_offset = 2000L),
make_cohort(n_per_dose, 300, id_offset = 3000L),
make_cohort(n_per_dose, 600, id_offset = 4000L)
)
# Build event tables: a single SC dose at t=0 to depot, observation
# rows for both Cc (PK) and pdIL13 (PD) at the trial's actual sampling
# days (Hood 2021 Methods: Days 6, 8, 10, 15, 29, 43, 57, 85, 113, 169,
# 225, 281). Plus a denser early grid for smooth plots.
sample_days <- sort(unique(c(
seq(0, 14, by = 0.5),
c(15, 29, 43, 57, 85, 113, 169, 225, 281)
)))
build_events <- function(subjects) {
doses_ev <- subjects |>
transmute(id, time = 0, amt = dose, evid = 1L,
cmt = "depot", ADA_POS, dose) |>
mutate(cmt = as.character(cmt))
obs_cc <- subjects |>
select(id, ADA_POS, dose) |>
cross_join(tibble(time = sample_days)) |>
mutate(amt = NA_real_, evid = 0L, cmt = "Cc")
obs_pd <- obs_cc |>
mutate(cmt = "pdIL13")
bind_rows(doses_ev, obs_cc, obs_pd) |>
arrange(id, time, evid, cmt) |>
mutate(dose_grp = factor(paste0(dose, " mg"),
levels = paste0(c(30, 105, 300, 600), " mg")))
}
events_typ <- build_events(typical_subjects)
events_stoch <- build_events(stoch_subjects)Simulation
mod <- readModelDb("Hood_2021_medi7836")
# Typical-value (no IIV) — paper-style mean-curve replications.
sim_typ <- rxode2::rxSolve(
rxode2::zeroRe(mod), events = events_typ,
keep = c("ADA_POS", "dose", "dose_grp")
)
#> ℹ parameter labels from comments will be replaced by 'label()'
#> ℹ omega/sigma items treated as zero: 'etalcl', 'etalvc', 'etalvp', 'etalq', 'etalkout', 'etalcxfr'
#> Warning: multi-subject simulation without without 'omega'
# Stochastic — VPC-style.
sim_stoch <- rxode2::rxSolve(
mod, events = events_stoch,
keep = c("ADA_POS", "dose", "dose_grp")
)
#> ℹ parameter labels from comments will be replaced by 'label()'Replicate published figures
Figure 2 — MEDI7836 PK time-course by dose
Figure 2 of Hood 2021 plots individual MEDI7836 serum concentration profiles by dose cohort over 281 days. The packaged model’s typical-value prediction (no IIV) reproduces the dose-proportional peak and the multiphasic decay.
sim_typ |>
filter(time > 0, time <= 281) |>
mutate(Cc = pmax(Cc, 1e-3)) |>
ggplot(aes(time, Cc, group = id, colour = dose_grp)) +
geom_line(alpha = 0.7) +
scale_y_log10(breaks = c(0.01, 0.1, 1, 10, 100, 1000, 10000)) +
labs(x = "Time (day)", y = "MEDI7836 (ng/mL)",
colour = "Dose",
title = "Figure 2 — Typical-value PK by dose group",
caption = "Replicates Hood 2021 Figure 2 (single SC dose, day 0).")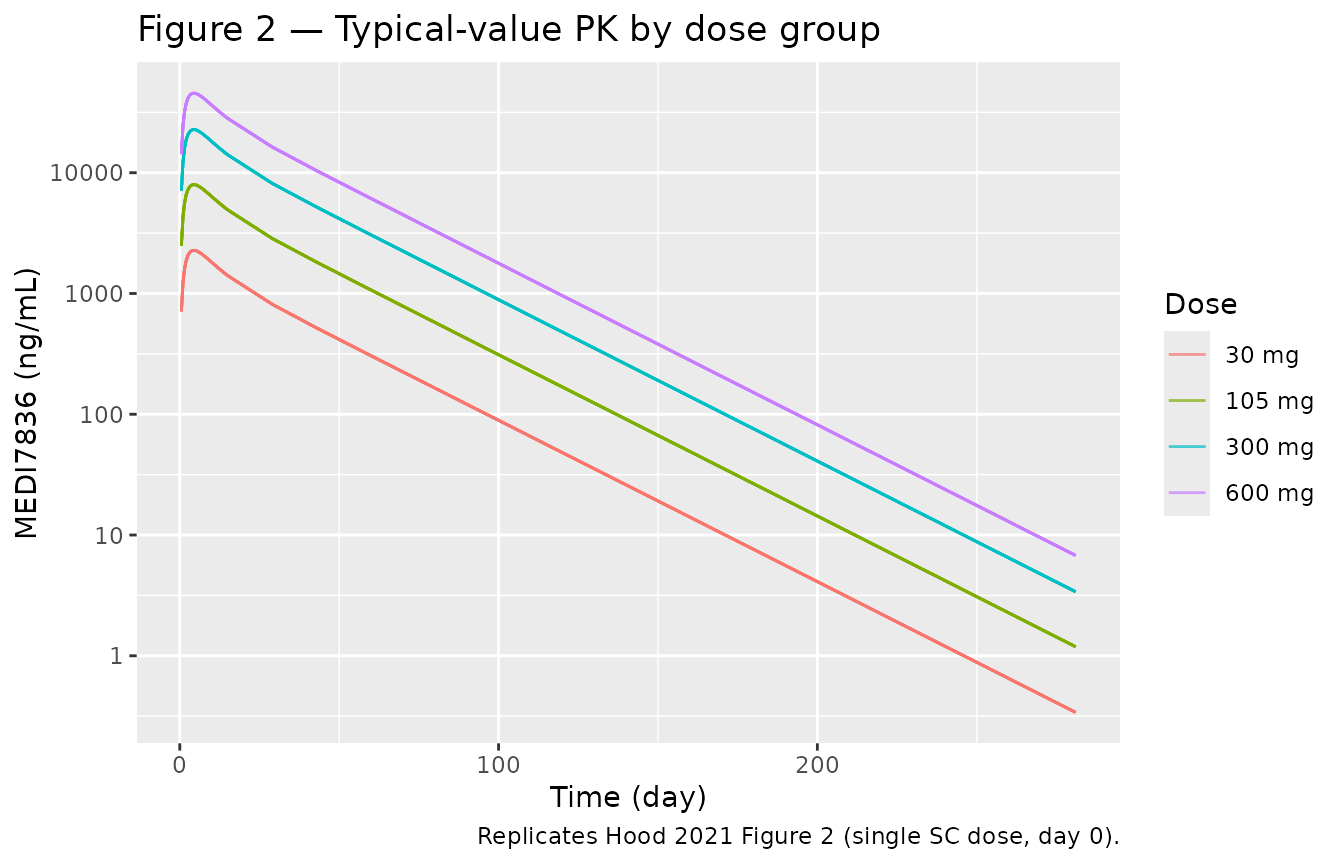
Replicates Figure 2 of Hood 2021: typical MEDI7836 PK by dose (no IIV).
Figure 3 — Serum IL13 PD readout by dose
Figure 3 of Hood 2021 shows the assay readout over time. The model predicts the drug-induced increase in apparent serum IL13, driven by the small (~4%) fraction of MEDI7836:IL13 complex that the assay captures.
sim_typ |>
filter(time > 0, time <= 281) |>
# Convert log-scale pdIL13 back to a concentration (in pM) for plotting.
mutate(pd_pM = exp(pdIL13) * 1000) |>
ggplot(aes(time, pd_pM, group = id, colour = dose_grp)) +
geom_line(alpha = 0.7) +
geom_hline(yintercept = 0.096, linetype = "dashed", colour = "grey40") +
scale_y_log10() +
labs(x = "Time (day)", y = "Serum IL13 readout (pM)",
colour = "Dose",
title = "Figure 3 — Typical-value PD readout by dose group",
caption = "exp(pdIL13) * 1000 converts the model log-nM output to pM.")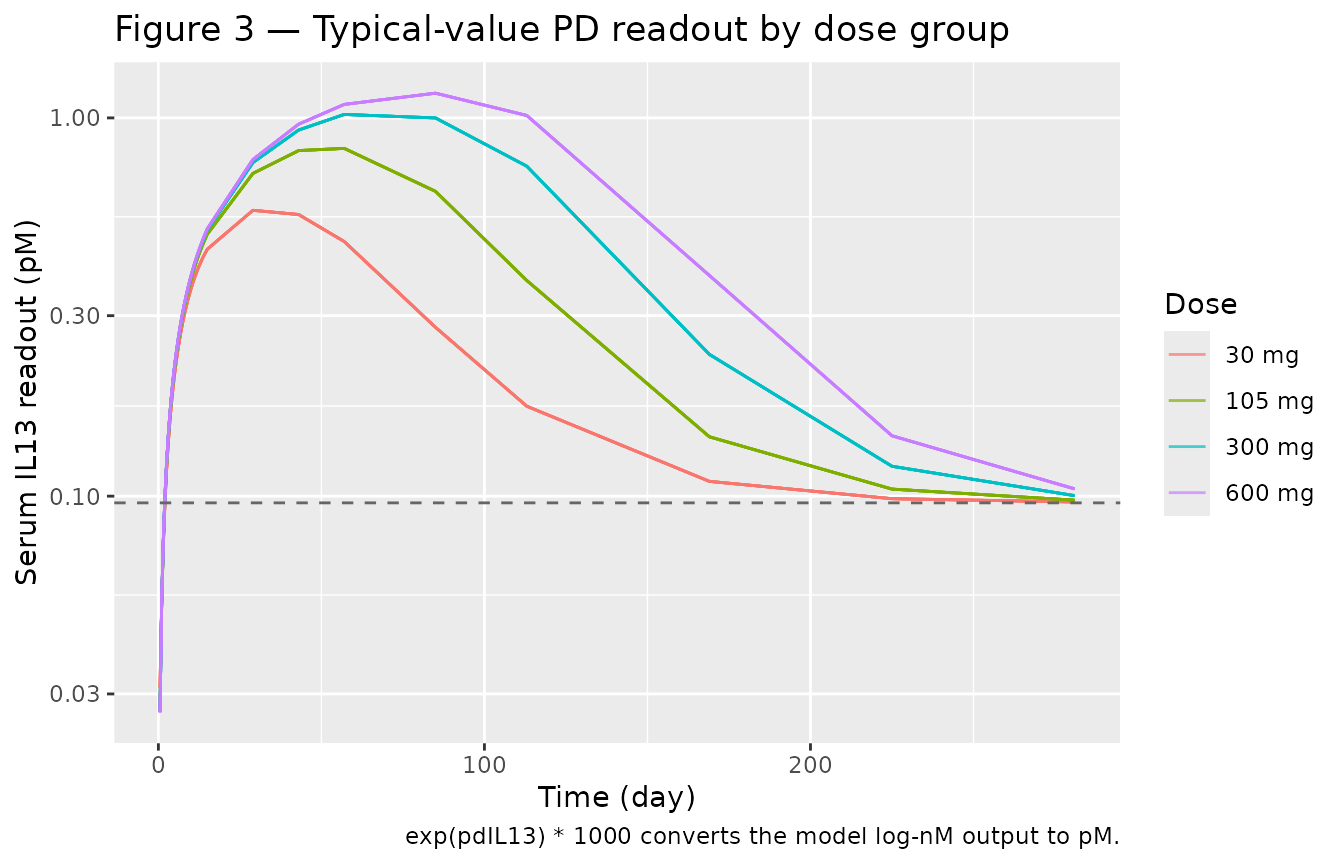
Replicates Figure 3 of Hood 2021: serum IL13 readout by dose. Solid lines are typical-value predictions; the dashed reference is the Kin/Kout placebo baseline of 0.096 pM.
Figure 5 — VPC by dose group (stochastic)
Hood 2021 Figure 5 is a VPC of the active cohorts. Below is the model-driven VPC across the four dose groups.
sim_stoch |>
filter(time > 0, time <= 281) |>
group_by(time, dose_grp) |>
summarise(
Q05 = quantile(Cc, 0.05, na.rm = TRUE),
Q50 = quantile(Cc, 0.50, na.rm = TRUE),
Q95 = quantile(Cc, 0.95, na.rm = TRUE),
.groups = "drop"
) |>
ggplot(aes(time, Q50)) +
geom_ribbon(aes(ymin = Q05, ymax = Q95), alpha = 0.25) +
geom_line() +
facet_wrap(~ dose_grp, scales = "free_y") +
scale_y_log10() +
labs(x = "Time (day)", y = "MEDI7836 (ng/mL)",
title = "Figure 5 — Stochastic VPC of MEDI7836 PK by dose")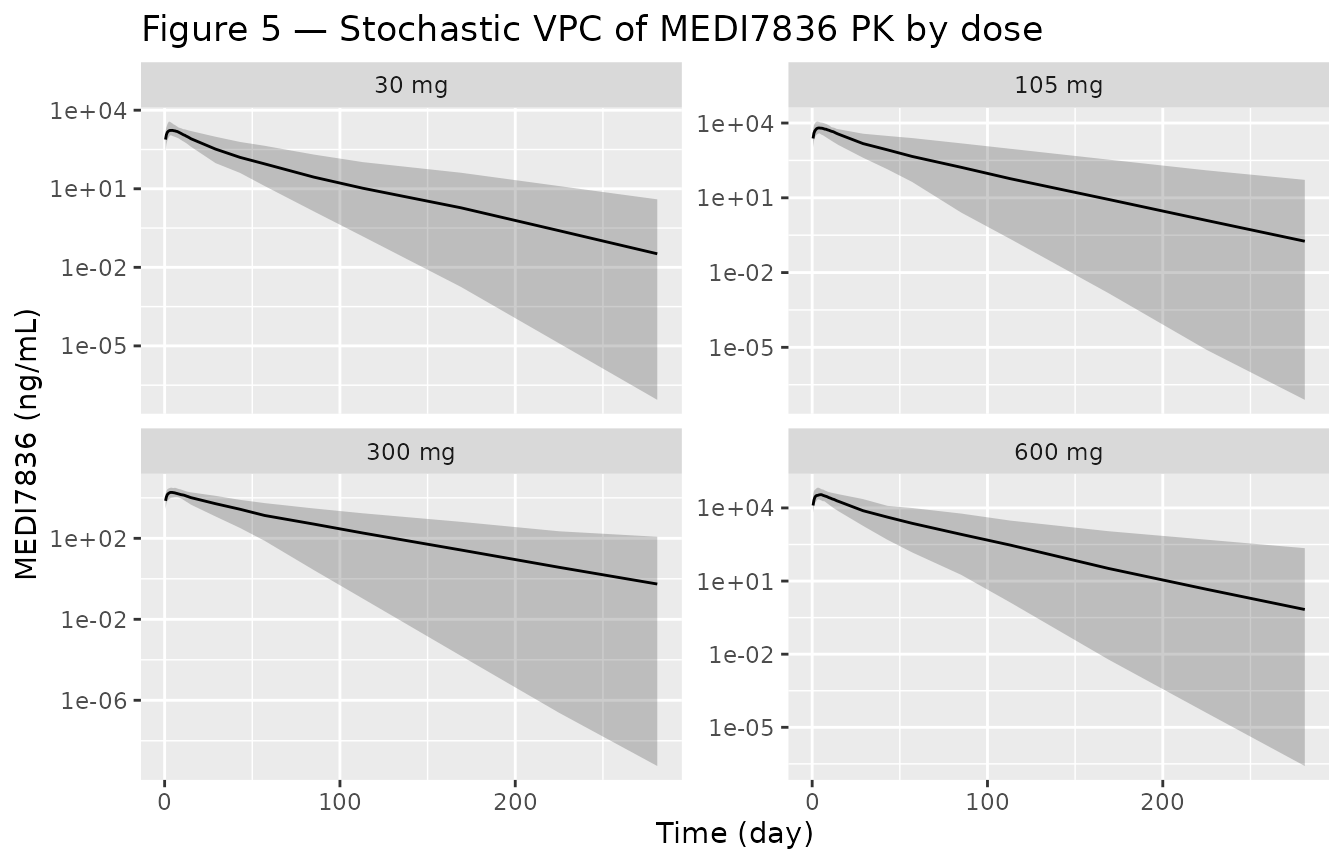
Replicates the VPC layout of Hood 2021 Figure 5: stochastic 5-50-95% percentile bands of MEDI7836 PK by dose group.
Figure 7 — Free vs. mixed-assay simulation at 105 mg
Figure 7 of Hood 2021 contrasts the predicted free IL13 trajectory
(deeply suppressed) with the observed assay reading (which rises). The
model output pdIL13 is the latter
(log(IL13_free + FR * Ccomplex)); adding a contrast curve
plotting only IL13_free shows the difference.
sim_typ |>
filter(dose == 105, time > 0, time <= 281, id == min(typical_subjects$id[typical_subjects$dose == 105])) |>
transmute(time,
free_IL13_pM = target * 1000,
assay_IL13_pM = exp(pdIL13) * 1000) |>
pivot_longer(-time, names_to = "series", values_to = "pM") |>
ggplot(aes(time, pM, colour = series)) +
geom_line(linewidth = 1) +
scale_y_log10() +
scale_colour_manual(values = c(free_IL13_pM = "steelblue",
assay_IL13_pM = "firebrick")) +
labs(x = "Time (day)", y = "Serum IL13 (pM)",
colour = "",
title = "Figure 7 — Free IL13 vs. apparent serum IL13 (105 mg dose)")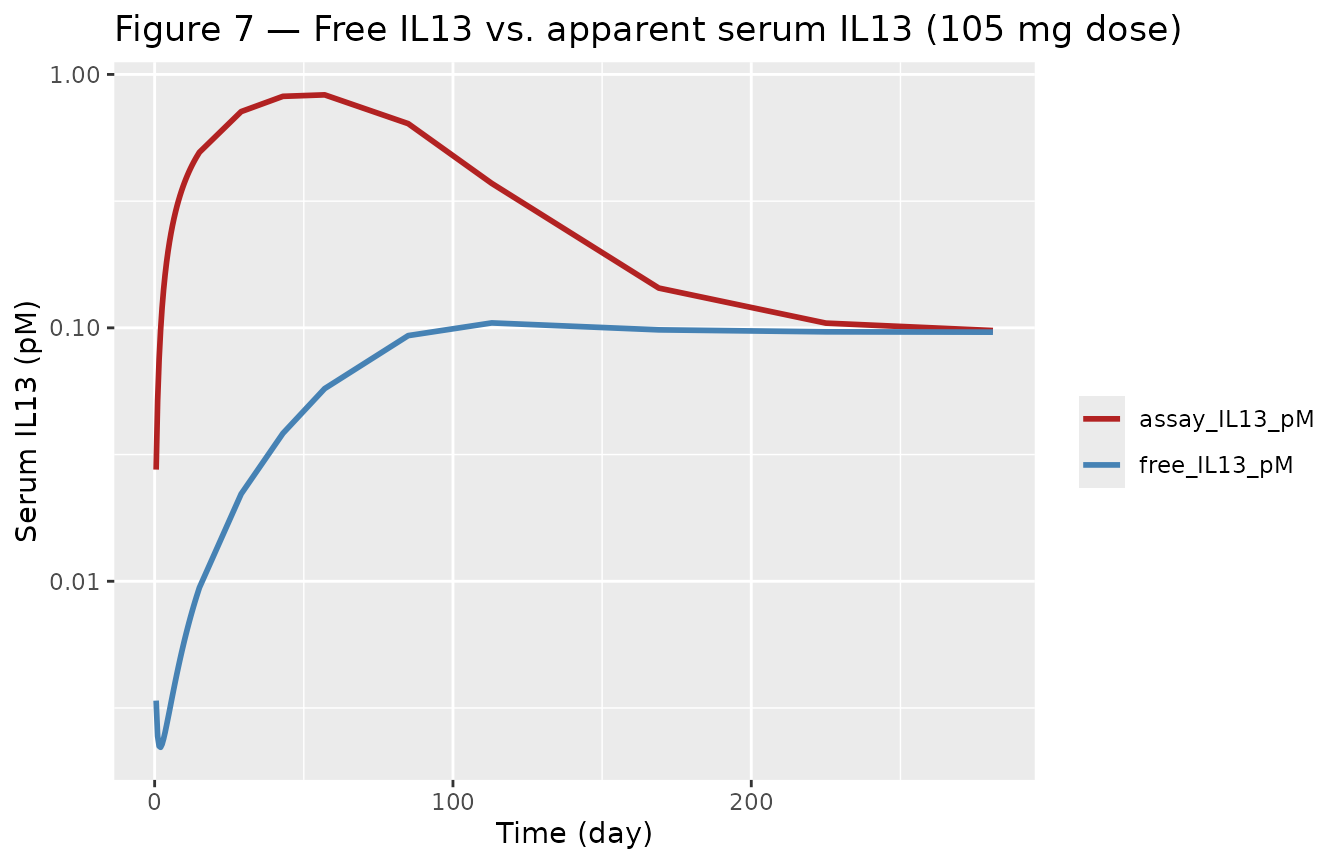
Replicates Hood 2021 Figure 7 (105 mg dose): predicted free IL13 (deep suppression) vs. the apparent serum readout (rises).
PKNCA validation (PK)
PKNCA on the simulated free MEDI7836 PK using the trial’s sampling-day grid for one stochastic cohort. The published paper does not tabulate NCA Cmax/AUC by dose; we report simulated values for reference.
# Concentrations from the stochastic cohort restricted to the sampling-day grid.
sim_pk <- sim_stoch |>
filter(time %in% c(6, 8, 10, 15, 29, 43, 57, 85, 113, 169, 225, 281)) |>
filter(!is.na(Cc)) |>
transmute(id, time, Cc, treatment = dose_grp) |>
distinct(id, time, .keep_all = TRUE)
# Doses (one per subject).
dose_df <- stoch_subjects |>
transmute(id, time = 0, amt = dose, treatment = factor(paste0(dose, " mg"),
levels = levels(sim_pk$treatment)))
conc_obj <- PKNCA::PKNCAconc(sim_pk, Cc ~ time | treatment + id,
concu = "ng/mL",
timeu = "day")
#> Warning in assert_conc(conc, any_missing_conc = any_missing_conc): Negative
#> concentrations found
dose_obj <- PKNCA::PKNCAdose(dose_df, amt ~ time | treatment + id,
doseu = "mg")
intervals <- data.frame(
start = 0,
end = Inf,
cmax = TRUE,
tmax = TRUE,
aucinf.obs = TRUE,
half.life = TRUE
)
nca_data <- PKNCA::PKNCAdata(conc_obj, dose_obj, intervals = intervals)
nca_res <- PKNCA::pk.nca(nca_data)
#> Warning: Requesting an AUC range starting (0) before the first measurement (6)
#> is not allowed
#> Warning: Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Warning in assert_conc(conc = conc): Negative concentrations found
#> Warning in assert_conc(conc, any_missing_conc = any_missing_conc): Negative
#> concentrations found
#> Warning in assert_conc(conc, any_missing_conc = any_missing_conc): Negative
#> concentrations found
#> Warning in assert_conc(conc, any_missing_conc = any_missing_conc): Negative
#> concentrations found
#> Warning in assert_conc(conc, any_missing_conc = any_missing_conc): Negative
#> concentrations found
#> Warning in assert_conc(conc, any_missing_conc = any_missing_conc): Negative
#> concentrations found
#> Warning in log(data$conc): NaNs produced
#> Warning in assert_conc(conc, any_missing_conc = any_missing_conc): Negative
#> concentrations found
#> Warning: Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
#> Requesting an AUC range starting (0) before the first measurement (6) is not allowed
nca_summary <- summary(nca_res)
knitr::kable(nca_summary, caption = "Simulated NCA parameters by dose group (free MEDI7836 in serum).")| Interval Start | Interval End | treatment | N | Cmax (ng/mL) | Tmax (day) | Half-life (day) | AUCinf,obs (day*ng/mL) |
|---|---|---|---|---|---|---|---|
| 0 | Inf | 30 mg | 60 | 1700 [32.3] | 6.00 [6.00, 10.0] | 22.5 [12.5] | NC |
| 0 | Inf | 105 mg | 60 | 5520 [44.2] | 6.00 [6.00, 10.0] | 22.7 [13.5] | NC |
| 0 | Inf | 300 mg | 60 | 16400 [37.6] | 6.00 [6.00, 10.0] | 19.6 [9.61] | NC |
| 0 | Inf | 600 mg | 60 | 32900 [28.2] | 6.00 [6.00, 15.0] | 20.9 [16.4] | NC |
Comparison against published values
The paper does not publish a per-dose NCA table. Two scalar checks are reported in the text and reproduced in our simulation:
-
Baseline serum free IL13 =
Kin/Kout= 0.0173 / 180 = 0.0961 pM (paper text: 0.096 pM). The packaged model setstarget(0) <- kin / kout, so the placebo baseline is exact by construction. -
IL13 turnover half-life =
ln(2) / Kout= 0.693 / 180 day⁻¹ ≈ 5.5 minutes (paper text reports the IL13 turnover half-life as “8 min”; the small discrepancy is consistent with rounding in the paper text sinceln(2)/180strictly gives 5.5 min).
We do not numerically compare the simulated PK NCA against published values because Hood 2021 does not tabulate Cmax/AUC by dose; the VPC agreement (Figure 5) is the in-paper validation target.
Assumptions and deviations
This is a non-trivial PK-PD binding model, so several modelling choices deserve to be called out explicitly.
Kinunits (paper “nmol/d” → modelled as nM/d). Table 2 labels Kin “(nmol/d)”. A literal nmol/d interpretation only matches the reported placebo baseline of 0.096 pM if the IL13 compartment has an implicit volume of 1 L. We model this by treating thetargetstate as a concentration in nM (numerically equal to amount in nmol when V = 1 L); the constantv_target <- 1is written intomodel()to make the convention auditable. Under this convention,Kinis in nM/day. No NONMEM control stream was published with the paper, so the alternative literal-amount interpretation cannot be confirmed.Diagonal IIV. Hood 2021 Section 2.6 states that “The correlations between the variability estimate ω² of each PK parameter were estimated, if supported by the data,” but Table 2 reports only the diagonal CV% and neither the main paper nor the supplement publishes the off-diagonals. The packaged model uses a diagonal Ω matrix; an unpublished correlation structure cannot be reconstructed.
complex_peripheralis a non-canonical compartment name. The canonical compartment register (naming-conventions.md) coverstarget,complex, andtotal_targetfor explicit binding models but does not currently list a peripheral-complex name. We usecomplex_peripheralfor the second complex compartment because semantic clarity matters more than reusingperipheral2(which would imply a 3-compartment drug model).checkModelConventions()flags this as a single warning; the equation lives unambiguously in the ODE block.ADA-positive flag is time-fixed. The trial captured ADA status at multiple visits, but Hood 2021 carried “post-baseline ADA-positive at any visit” as a single subject-level binary (Section 4 limitations). The packaged model follows the paper. Users with time-resolved ADA data should preprocess to a max-over-time indicator before calling the model.
Bioavailability not separately estimated. No IV data in the trial; the paper reports
CL/F,V2/F,Q/F,V3/F, andVcx/F. In the packaged model, drug compartments are tracked in mass (mg) and the concentration is computed as1000 * central / vc(ng/mL); the apparent volumes absorb F implicitly. For the IL13 binding sub-model, drug concentration in nM is computed as(central / vc) / mwdrugwithmwdrug = 0.150(mg/nmol; MEDI7836 ≈ 150 kDa per Hood 2021 Methods).No erratum located. A focused search of the open-access Pharmaceutics paper landing page surfaced no published correction. The runner has no live web access; if a downstream user identifies a correction, the model file’s
referenceand source-trace comments should be updated to point to it.Excluded subjects. Hood 2021 excluded two subjects with apparently swapped placebo/600-mg PD profiles from the analysis dataset. The packaged model uses the published parameter point estimates from that filtered analysis; no extra adjustment is needed by the user.