
Single Subject ODE solving -- differences from multiple subject
2026-05-25
Source:vignettes/rxode2-single-subject.Rmd
rxode2-single-subject.RmdSingle Subject solving
Originally, rxode2 was only created to solve ODEs for one individual. That is a single system without any changes in individual parameters.
Of course this is still supported, the classic examples are found in rxode2 intro.
This article discusses the differences between multiple subject and single subject solving. There are three differences:
- Single solving does not solve each ID in parallel
- Single solving lacks the
idcolumn in parameters($params) as well as in the actual dataset. - Single solving allows parameter exploration easier because each parameter can be modified. With multiple subject solves, you have to make sure to update each individual parameter.
The first obvious difference is in speed; With multiple subjects you can run each subject ID in parallel. For more information and examples of the speed gains with multiple subject solving see the Speeding up rxode2 vignette.
The next difference is the amount of information output in the final data.
Taking the 2 compartment indirect response model originally in the tutorial:
library(rxode2)
#> rxode2 5.1.0 using 2 threads (see ?getRxThreads)
#> no cache: create with `rxCreateCache()`
mod1 <- function() {
ini({
KA=2.94E-01
CL=1.86E+01
V2=4.02E+01
Q=1.05E+01
V3=2.97E+02
Kin=1
Kout=1
EC50=200
})
model({
C2 = centr/V2
C3 = peri/V3
d/dt(depot) =-KA*depot
d/dt(centr) = KA*depot - CL*C2 - Q*C2 + Q*C3
d/dt(peri) = Q*C2 - Q*C3
d/dt(eff) = Kin - Kout*(1-C2/(EC50+C2))*eff
eff(0) = 1
})
}
et <- et(amount.units='mg', time.units='hours') |>
et(dose=10000, addl=9, ii=12) |>
et(amt=20000, nbr.doses=5, start.time=120, dosing.interval=24) |>
et(0:240) # samplingNow a simple solve
x <- rxSolve(mod1, et)
#> ℹ parameter labels from comments are typically ignored in non-interactive mode
#> ℹ Need to run with the source intact to parse comments
x
#> ── Solved rxode2 object ──
#> ── Parameters (x$params): ──
#> KA CL V2 Q V3 Kin Kout EC50
#> 0.294 18.600 40.200 10.500 297.000 1.000 1.000 200.000
#> ── Initial Conditions (x$inits): ──
#> depot centr peri eff
#> 0 0 0 1
#> ── First part of data (object): ──
#> # A tibble: 241 × 7
#> time C2 C3 depot centr peri eff
#> [h] <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 0 0 0 30000 0 0 1
#> 2 1 133. 2.76 22358. 5352. 820. 1.21
#> 3 2 165. 8.02 16663. 6619. 2382. 1.46
#> 4 3 156. 13.4 12419. 6260. 3971. 1.61
#> 5 4 133. 17.9 9255. 5366. 5329. 1.66
#> 6 5 109. 21.5 6898. 4400. 6395. 1.63
#> # ℹ 235 more rows
print(x)
#> ── Solved rxode2 object ──
#> ── Parameters ($params): ──
#> KA CL V2 Q V3 Kin Kout EC50
#> 0.294 18.600 40.200 10.500 297.000 1.000 1.000 200.000
#> ── Initial Conditions ($inits): ──
#> depot centr peri eff
#> 0 0 0 1
#> ── First part of data (object): ──
#> # A tibble: 241 × 7
#> time C2 C3 depot centr peri eff
#> [h] <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 0 0 0 30000 0 0 1
#> 2 1 133. 2.76 22358. 5352. 820. 1.21
#> 3 2 165. 8.02 16663. 6619. 2382. 1.46
#> 4 3 156. 13.4 12419. 6260. 3971. 1.61
#> 5 4 133. 17.9 9255. 5366. 5329. 1.66
#> 6 5 109. 21.5 6898. 4400. 6395. 1.63
#> # ℹ 235 more rows
plot(x, C2, eff)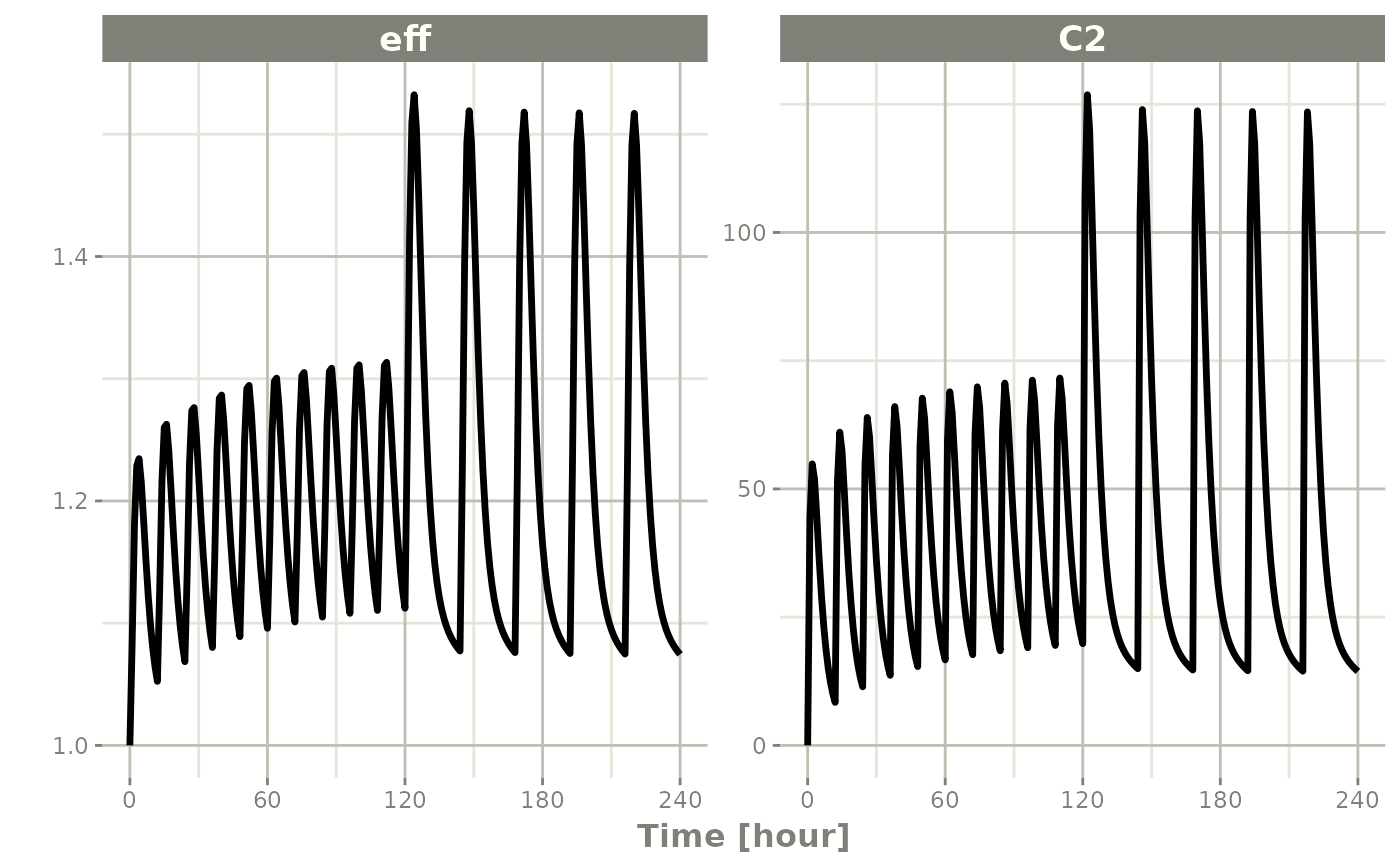
To better see the differences between the single solve, you can solve for 2 individuals
x2 <- rxSolve(mod1, et |> et(id=1:2), params=data.frame(CL=c(18.6, 7.6)))
#> ℹ parameter labels from comments are typically ignored in non-interactive mode
#> ℹ Need to run with the source intact to parse comments
print(x2)
#> ── Solved rxode2 object ──
#> ── Parameters ($params): ──
#> # A tibble: 2 × 9
#> id KA CL V2 Q V3 Kin Kout EC50
#> <fct> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 1 0.294 18.6 40.2 10.5 297 1 1 200
#> 2 2 0.294 7.6 40.2 10.5 297 1 1 200
#> ── Initial Conditions ($inits): ──
#> depot centr peri eff
#> 0 0 0 1
#> ── First part of data (object): ──
#> # A tibble: 482 × 8
#> id time C2 C3 depot centr peri eff
#> <int> [h] <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 1 0 0 0 30000 0 0 1
#> 2 1 1 133. 2.76 22358. 5352. 820. 1.21
#> 3 1 2 165. 8.02 16663. 6619. 2382. 1.46
#> 4 1 3 156. 13.4 12419. 6260. 3971. 1.61
#> 5 1 4 133. 17.9 9255. 5366. 5329. 1.66
#> 6 1 5 109. 21.5 6898. 4400. 6395. 1.63
#> # ℹ 476 more rows
plot(x2, C2, eff)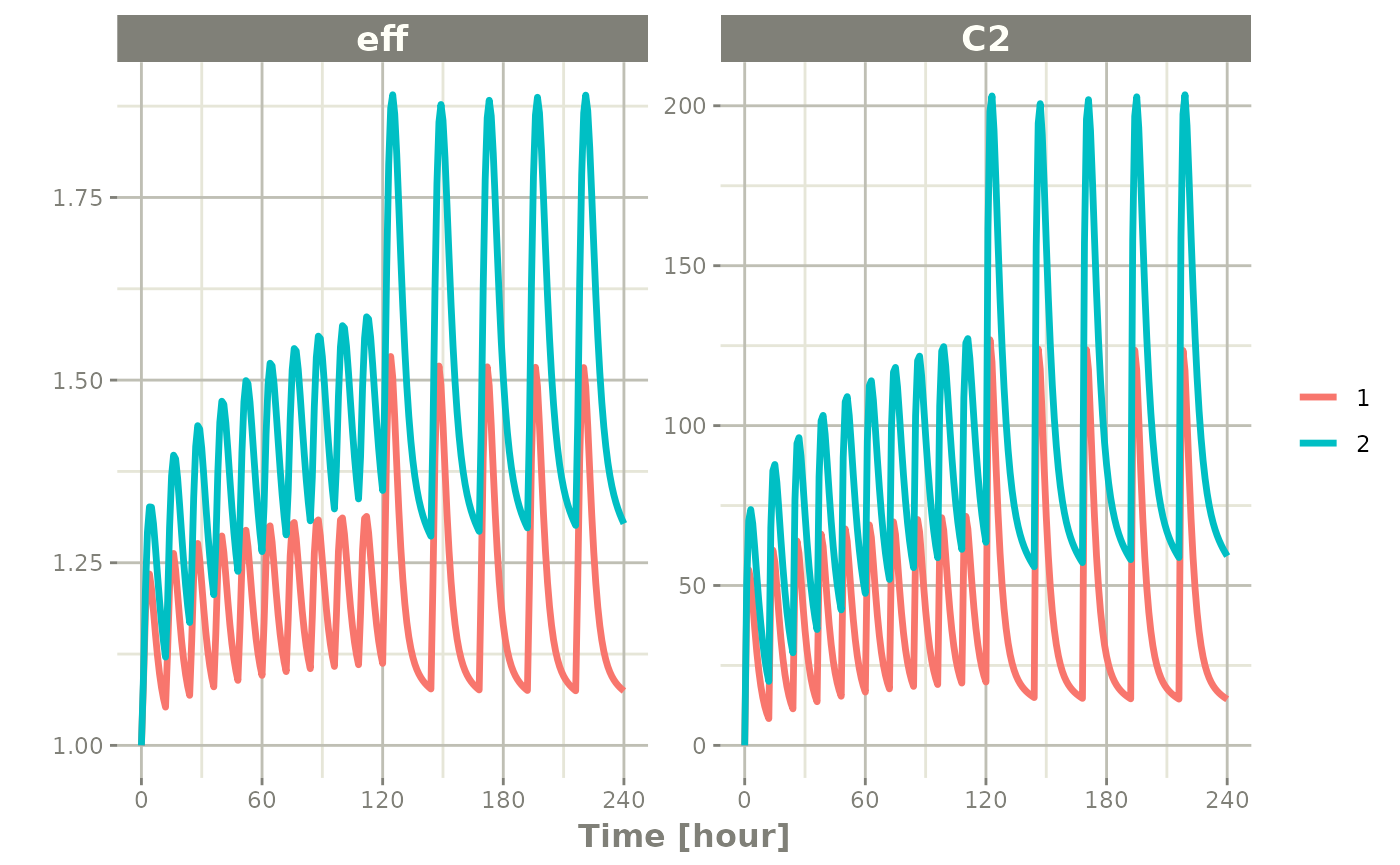
By observing the two solves, you can see:
- A multiple subject solve contains the
idcolumn both in the data frame and then data frame of parameters for each subject.
The last feature that is not as obvious, modifying the individual parameters. For single subject data, you can modify the rxode2 data frame changing initial conditions and parameter values as if they were part of the data frame, as described in the rxode2 Data Frames.
For multiple subject solving, this feature still works, but requires care when supplying each individual’s parameter value, otherwise you may change the solve and drop parameter for key individuals.
Summary of Single solve vs Multiple subject solving
| Feature | Single Subject Solve | Multiple Subject Solve |
|---|---|---|
| Parallel | None | Each Subject |
| $params | data.frame with one parameter value | data.frame with one parameter per subject (w/ID column) |
| solved data | Can modify individual parameters with $ syntax | Have to modify all the parameters to update solved object |